- 4192 Devonwood Way, Ashburn, Virginia, 20148
- Helpline: +1 (703) 665-3747
The Ploc_Bal-Mgpos is a Powerful Artificial Intelligence Tool for Predicting the Subcellular Localization of Gram-Positive Bacterial Proteins According to Their Sequence Information Alone
- Home
- Back to Journal
- Article Details
Recently a very useful web-server, or AI (Artificial Intelligence) tool, has been established for predicting the subcellular localization of Gram-positive bacterial proteins purely according to their sequences information for the multi-label systems [1], in which a same protein may occur or travel between two or more locations and hence its ID (identification) needs two or more labels as well, namely the “multi-label mark” [2].
The AI tool is named as “pLoc_bal-mGpos”, where “bal” stands for that the AI tool has been treated by balancing out the training dataset [3–9], and “m” for that the AI tool bears the capacity to deal with the multi-label systems. Below, let us demonstrate how the AI tool is working.
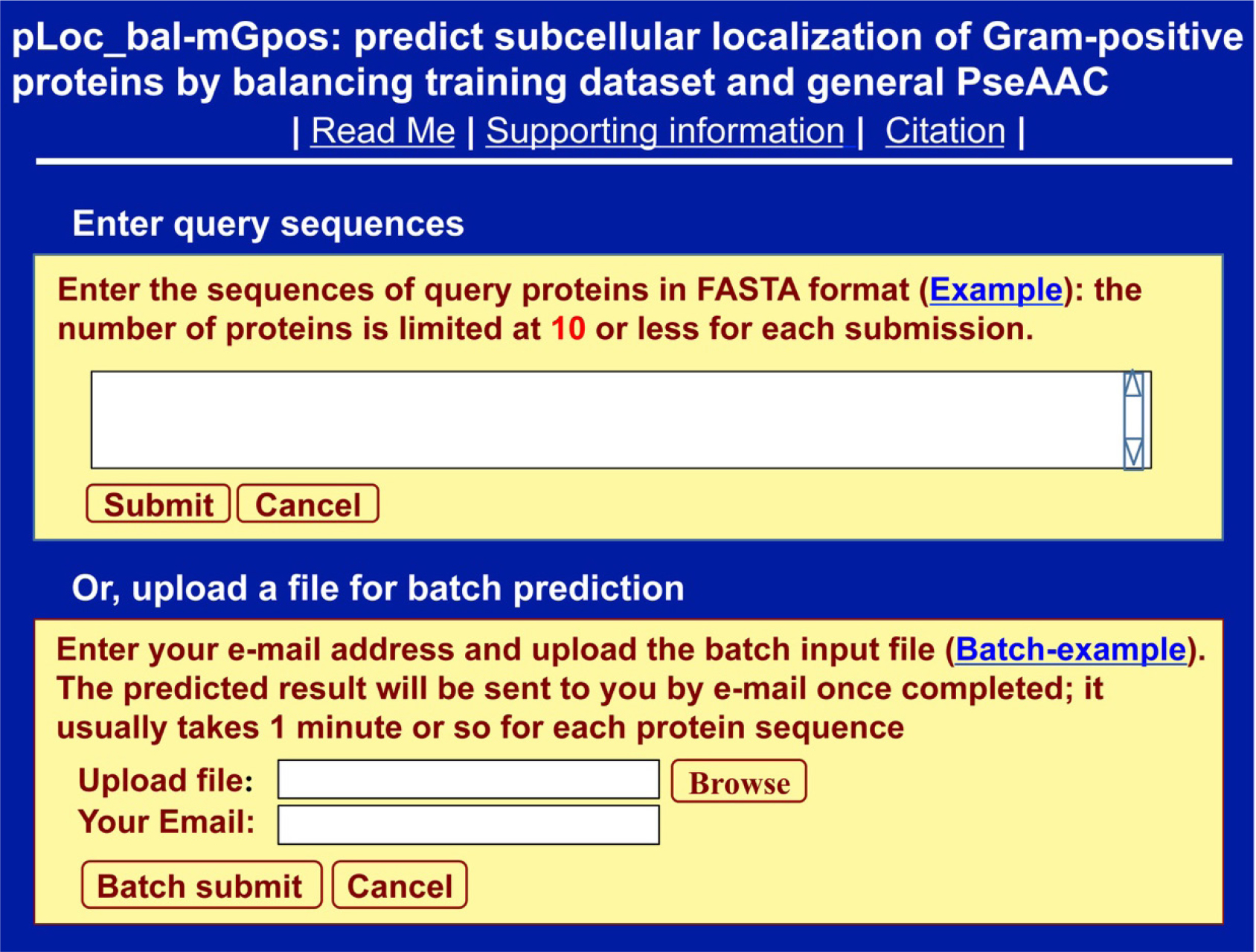
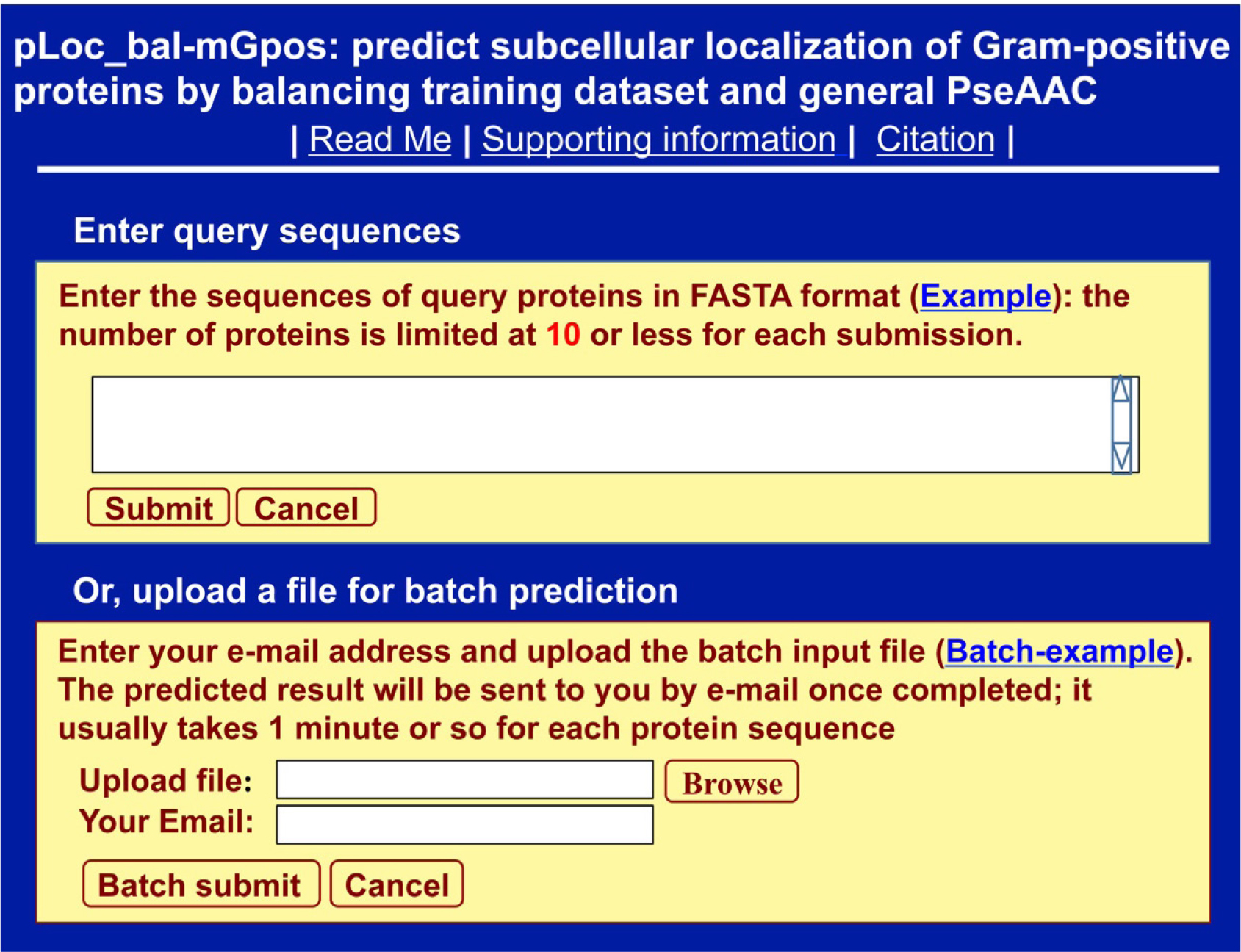
Clicking the link at http://www.jci-bioinfo.cn/pLoc_bal-mGpos/, you will see the top page of the pLoc_bal-mGpos web-server prompted on your computer’s screen (Figure 1). Then, click the Example button and use the query protein sequences as the input. After clicking the Submit button, you will see Figure 2 shown on the screen of your computer. The corresponding outcomes were detailed in [4]. You can see from there: nearly all the success rates achieved by the AI tool for the Gram-positive bacterial proteins in each of the 6 subcellular locations are within the range of 98–99%. Such a high prediction quality is far beyond the reach of any of its counterparts.
Figure 1. A semi screenshot for the top page of pLoc_bal-mGpos (Adapted from [4] with permission).
Figure 2. A semi screenshot for the webpage obtained by following Step 3 of Section 3.5 (Adapted from [4] with permission).
In addition to the advantages of high accuracy and easy to use, the AI tool has been constructed by strictly complying with the “Chou’s 5-steps rule” and hence possesses the following terrific merits as concurred by many investigators (see, e.g., [10–91] as well as three comprehensive review papers [2, 92, 93]): (1) crystal clear in logic development, (2) completely transparent in operation, (3) easily to repeat the reported results by other investigators, (4) with high potential in stimulating other sequence-analyzing methods, and (5) very convenient to be used by the majority of experimental scientists.
Besides, the approach [94–96] of PseAAC (Pseudo Amino Acid Composition) has also been used during the development of the AI tool. It is a very powerful approach for formulating the samples of proteins by catching their special features, as done by many investigators [97–222].
Moreover, the IHTS (Inserting Hypothetical Training Samples) treatment has also been utilized to balance out the training dataset [57, 60, 84].
For the wonderful and awesome roles of the “5-steps rule” in driving proteome, genome analyses and drug development, see a series of recent papers [2, 93, 223–233] where the rule and its wide applications have been very impressively presented from various aspects or at different angles.
References
- K.C. Chou, H.B. Shen, Recent progresses in protein subcellular location prediction. Analytical Biochemistry 370 (2007) 1–16.
- K.C. Chou, Advance in predicting subcellular localization of multi-label proteins and its implication for developing multi-target drugs. Current Medicinal Chemistry 26 (2019) 4918–4943.
- X. Xiao, X. Cheng, G. Chen, Q. Mao, K.C. Chou, pLoc_bal-mVirus: Predict Subcellular Localization of Multi-Label Virus Proteins by Chou’s General PseAAC and IHTS Treatment to Balance Training Dataset. Med Chem 15 (2019) 496–509.
- X. Xiao, X. Cheng, G. Chen, Q. Mao, K.C. Chou, pLoc_bal-mGpos: predict subcellular localization of Gram-positive bacterial proteins by quasi-balancing training dataset and PseAAC. Genomics 111 (2019) 886–892.
- K.C. Chou, X. Cheng, X. Xiao, pLoc_bal-mEuk: predict subcellular localization of eukaryotic proteins by general PseAAC and quasi-balancing training dataset. Med Chem 15 (2019) 472–485.
- K.C. Chou, X. Cheng, X. Xiao, pLoc_bal-mHum: predict subcellular localization of human proteins by PseAAC and quasi-balancing training dataset Genomics 111 (2019) 1274–1282.
- X. Cheng, W.Z. Lin, X. Xiao, K.C. Chou, pLoc_bal-mAnimal: predict subcellular localization of animal proteins by balancing training dataset and PseAAC. Bioinformatics 35 (2019) 398–406.
- X. Cheng, X. Xiao, K.C. Chou, pLoc_bal-mPlant: predict subcellular localization of plant proteins by general PseAAC and balancing training dataset Curr Pharm Des 24 (2018) 4013–4022.
- X. Cheng, X. Xiao, K.C. Chou, pLoc_bal-mGneg: predict subcellular localization of Gram-negative bacterial proteins by quasi-balancing training dataset and general PseAAC. Journal of Theoretical Biology 458 (2018) 92–102.
- A.H. Butt, Y.D. Khan, Prediction of S-Sulfenylation Sites Using Statistical Moments Based Features via Chou’s 5-Step Rule. International Journal of Peptide Research and Therapeutics (IJPRT) https://doi.org/10.1007/s10989-019-09931-2 (2018).
- M. Awais, W. Hussain, Y.D. Khan, N. Rasool, S.A. Khan, K.C. Chou, iPhosH-PseAAC: Identify phosphohistidine sites in proteins by blending statistical moments and position relative features according to the Chou’s 5-step rule and general pseudo amino acid composition. IEEE/ACM Trans Comput Biol Bioinform https://doi.org/10.1109/TCBB.2019.2919025 or https://www.ncbi.nlm.nih.gov/pubmed/31144645 (2019).
- O. Barukab, Y.D. Khan, S.A. Khan, K.C. Chou, iSulfoTyr-PseAAC: Identify tyrosine sulfation sites by incorporating statistical moments via Chou’s 5-steps rule and pseudo components Current Genomics https:/doi.org/10.2174/1389202920666190819091609 or http://www.eurekaselect.com/174277/article (2019).
- A.H. Butt, Y.D. Khan, Prediction of S-Sulfenylation Sites Using Statistical Moments Based Features via Chou’s 5-Step Rule. International Journal of Peptide Research and Therapeutics (IJPRT) htpps://doi.org/10.1007/s10989-019-09931-2 (2019).
- Y. Chen, X. Fan, Use of Chou’s 5-Steps Rule to Reveal Active Compound and Mechanism of Shuangshen Pingfei San on Idiopathic Pulmonary Fibrosis. Current Molecular Medicine https://doi.org/10.2174/1566524019666191011160543 (2019).
- X. Du, Y. Diao, H. Liu, S. Li, MsDBP: Exploring DNA-binding Proteins by Integrating Multi-scale Sequence Information via Chou’s 5-steps Rule. Journal of Proteome Research 18 (2019) 3119–3132.
- A. Ehsan, M.K. Mahmood, Y.D. Khan, O.M. Barukab, S.A. Khan, K.C. Chou, iHyd-PseAAC (EPSV): Identify hydroxylation sites in proteins by extracting enhanced position and sequence variant feature via Chou’s 5-step rule and general pseudo amino acid composition. Current Genomics 20 (2019) 124–133.
- W. Hussain, S.D. Khan, N. Rasool, S.A. Khan, K.C. Chou, SPalmitoylC-PseAAC: A sequence-based model developed via Chou’s 5-steps rule and general PseAAC for identifying S-palmitoylation sites in proteins. Anal Biochem 568 (2019) 14–23.
- W. Hussain, Y.D. Khan, N. Rasool, S.A. Khan, K.C. Chou, SPrenylC-PseAAC: A sequence-based model developed via Chou’s 5-steps rule and general PseAAC for identifying S-prenylation sites in proteins. J Theor Biol 468 (2019) 1–11.
- Z. Ju, S.Y. Wang, Prediction of lysine formylation sites using the composition of k-spaced amino acid pairs via Chou’s 5-steps rule and general pseudo components. Genomics htpps:/doi.org/10.1016/j.ygeno.2019.05.027 or https://www.ncbi.nlm.nih.gov/pubmed/31175975 (2019).
- M. Kabir, S. Ahmad, M. Iqbal, M. Hayat, iNR-2L: A two-level sequence-based predictor developed via Chou’s 5-steps rule and general PseAAC for identifying nuclear receptors and their families. Genomics https://www.ncbi.nlm.nih.gov/pubmed/30779939 (2019).
- Z.U. Khan, F. Ali, I.A. Khan, Y. Hussain, D. Pi, iRSpot-SPI: Deep learning-based recombination spots prediction byincorporating secondary sequence information coupled withphysio-chemical properties via Chou’s 5-step rule and pseudo components. Chemometrics and Intelligent Laboratory Systems (CHEMOLAB) 189 (2019) 169–180.
- J. Lan, J. Liu, C. Liao, D.J. Merkler, Q. Han, J. Li, A Study for Therapeutic Treatment against Parkinson’s Disease via Chou’s 5-steps Rule. Current Topics in Medicinal Chemistry https://doi.org/10.2174/1568026619666191019111528 or http://www.eurekaselect.com/175887/article (2019).
- N.Q.K. Le, iN6-methylat (5-step): identifying DNA N(6)-methyladenine sites in rice genome using continuous bag of nucleobases via Chou’s 5-step rule. Mol Genet Genomics 294 (2019) 1173–1182.
- N.Q.K. Le, E.K.Y. Yapp, Q.T. Ho, N. Nagasundaram, Y.Y. Ou, H.Y. Yeh, iEnhancer-5Step: Identifying enhancers using hidden information of DNA sequences via Chou’s 5-step rule and word embedding. Anal Biochem 571 (2019) 53–61.
- N.Q.K. Le, E.K.Y. Yapp, Y.Y. Ou, H.Y. Yeh, iMotor-CNN: Identifying molecular functions of cytoskeleton motor proteins using 2D convolutional neural network via Chou’s 5-step rule. Anal Biochem 575 (2019) 17–26.
- R. Liang, J. Xie, C. Zhang, M. Zhang, H. Huang, H. Huo, X. Cao, B. Niu, Identifying Cancer Targets Based on Machine Learning Methods via Chou’s 5-steps Rule and General Pseudo Components. Current Topics in Medicnal Chemistry htpps://doi.org/10.2174/1568026619666191016155543 (2019).
- Y. Liang, S. Zhang, Identifying DNase I hypersensitive sites using multi-features fusion and F-score features selection via Chou’s 5-steps rule. Biophys Chem 253 (2019) 106227.
- S.J. Malebary, M.S.U. Rehman, Y.D. Khan, iCrotoK-PseAAC: Identify lysine crotonylation sites by blending position relative statistical features according to the Chou’s 5-step rule. PLoS One 14 (2019) e0223993.
- I. Nazari, M. Tahir, H. Tayari, K.T. Chong, iN6-Methyl (5-step): Identifying RNA N6-methyladenosine sites using deep learning mode via Chou’s 5-step rules and Chou’s general PseKNC. Chemometrics and Intelligent Laboratory Systems (CHEMOLAB) htpps://doi.org/10.1016/j.chemolab.2019.103811 (2019).
- Q. Ning, Z. Ma, X. Zhao, dForml(KNN)-PseAAC: Detecting formylation sites from protein sequences using K-nearest neighbor algorithm via Chou’s 5-step rule and pseudo components. J Theor Biol 470 (2019) 43–49.
- Salman, M. Khan, N. Iqbal, T. Hussain, S. Afzal, K.C. Chou, A two-level computation model based on deep learning algorithm for identification of piRNA and their functions via Chou’s 5-steps rule. International Journal of Peptide Research and Therapeutics (IJPRT) htpps:/doi.org/10.1007/s10989-019-09887-3 (2019).
- M. Tahir, H. Tayara, K.T. Chong, iDNA6mA (5-step rule): Identification of DNA N6-methyladenine sites in the rice genome by intelligent computational model via Chou’s 5-step rule. CHEMOLAB 189 (2019) 96–101.
- S. Vishnoi, P. Garg, P. Arora, Physicochemical n-Grams Tool: A tool for protein physicochemical descriptor generation via Chou’s 5-steps rule. Chem Biol Drug Des htpps://doi.org/10.1111/cbdd.13617 or https://www.ncbi.nlm.nih.gov/pubmed/31483930 (2019).
- A. Wiktorowicz, A. Wit, A. Dziewierz, L. Rzeszutko, D. Dudek, P. Kleczynski, Calcium Pattern Assessment in Patients with Severe Aortic Stenosis Via the Chou’s 5-Steps Rule. Current Pharmaceutical Design https:/doi.org/10.2174/1381612825666190930101258 (2019).
- L. Yang, Y. Lv, S. Wang, Q. Zhang, Y. Pan, D. Su, Q. Lu, Y. Zuo, Identifying FL11 subtype by characterizing tumor immune microenvironment in prostate adenocarcinoma via Chou’s 5-steps rule. Genomics https://doi.org/10.1016/j.ygeno.2019.08.021 (2019).
- L. Yang, Y. Lv, S. Wang, Q. Zhang, Y. Pan, D. Su, Q. Lu, Y. Zuo, Identifying FL11 subtype by characterizing tumor immune microenvironment in prostate adenocarcinoma via Chou’s 5-steps rule. Genomics (2019).
- Y.D. Khan, N. Amin, W. Hussain, N. Rasool, S.A. Khan, K.C. Chou, iProtease-PseAAC(2L): A two-layer predictor for identifying proteases and their types using Chou’s 5-step-rule and general PseAAC. Anal Biochem 588 (2020) 113477.
- Y. Xu, J. Ding, L.Y. Wu, K.C. Chou, iSNO-PseAAC: Predict cysteine S-nitrosylation sites in proteins by incorporating position specific amino acid propensity into pseudo amino acid composition PLoS ONE 8 (2013) e55844.
- Y. Xu, X.J. Shao, L.Y. Wu, N.Y. Deng, K.C. Chou, iSNO-AAPair: incorporating amino acid pairwise coupling into PseAAC for predicting cysteine S-nitrosylation sites in proteins. PeerJ 1 (2013) e171.
- Y. Xu, X. Wen, X.J. Shao, N.Y. Deng, K.C. Chou, iHyd-PseAAC: Predicting hydroxyproline and hydroxylysine in proteins by incorporating dipeptide position-specific propensity into pseudo amino acid composition. International Journal of Molecular Sciences (IJMS) 15 (2014) 7594–7610.
- Y. Xu, X. Wen, L.S. Wen, L.Y. Wu, N.Y. Deng, K.C. Chou, iNitro-Tyr: Prediction of nitrotyrosine sites in proteins with general pseudo amino acid composition. PLoS ONE 9 (2014) e105018.
- Y. Xu, K.C. Chou, Recent progress in predicting posttranslational modification sites in proteins. Curr Top Med Chem 16 (2016) 591–603.
- L.M. Liu, Y. Xu, K.C. Chou, iPGK-PseAAC: identify lysine phosphoglycerylation sites in proteins by incorporating four different tiers of amino acid pairwise coupling information into the general PseAAC. Med Chem 13 (2017) 552–559.
- Y. Xu, C. Li, K.C. Chou, iPreny-PseAAC: identify C-terminal cysteine prenylation sites in proteins by incorporating two tiers of sequence couplings into PseAAC. Med Chem 13 (2017) 544–551.
- L. Cai, C.L. Wan, L. He, S. Jong, K.C. Chou, Gestational influenza increases the risk of psychosis in adults. Medicinal Chemistry 11 (2015) 676–682.
- J. Liu, J. Song, M.Y. Wang, L. He, L. Cai, K.C. Chou, Association of EGF rs4444903 and XPD rs13181 polymorphisms with cutaneous melanoma in Caucasians. Medicinal Chemistry 11 (2015) 551–559.
- L. Cai, Y.H. Yang, L. He, K.C. Chou, Modulation of cytokine network in the comorbidity of schizophrenia and tuberculosis. Curr Top Med Chem 16 (2016) 655–665.
- L. Cai, W. Yuan, Z. Zhang, L. He, K.C. Chou, In-depth comparison of somatic point mutation callers based on different tumor next-generation sequencing depth data Scientific Reports 6 (2016) 36540.
- Y. Zhu, Q.W. Cong, Y. Liu, C.L. Wan, T. Yu, G. He, L. He, L. Cai, K.C. Chou, Antithrombin is an importantly inhibitory role against blood clots. Curr Top Med Chem 16 (2016) 666–674.
- Z.D. Zhang, K. Liang, K. Li, G.Q. Wang, K.W. Zhang, L. Cai, S.T. Zha, K.C. Chou, Chlorella vulgaris induces apoptosis of human non-small cell lung carcinoma (NSCLC) cells. Med Chem 13 (2017) 560–568.
- L. Cai, T. Huang, J. Su, X. Zhang, W. Chen, F. Zhang, L. He, K.C. Chou, Implications of newly identified brain eQTL genes and their interactors in Schizophrenia. Molecular Therapy - Nucleic Acids 12 (2018) 433–442.
- B. Niu, M. Zhang, P. Du, L. Jiang, R. Qin, Q. Su, F. Chen, D. Du, Y. Shu, K.C. Chou, Small molecular floribundiquinone B derived from medicinal plants inhibits acetylcholinesterase activity. Oncotarget 8 (2017) 57149–57162.
- Q. Su, W. Lu, D. Du, F. Chen, B. Niu, K.C. Chou, Prediction of the aquatic toxicity of aromatic compounds to tetrahymena pyriformis through support vector regression. Oncotarget 8 (2017) 49359–49369.
- Y. Lu, S. Wang, J. Wang, G. Zhou, Q. Zhang, X. Zhou, B. Niu, Q. Chen, K.C. Chou, An Epidemic Avian Influenza Prediction Model Based on Google Trends. Letters in Organic Chemistry 16 (2019) 303–310.
- B. Niu, C. Liang, Y. Lu, M. Zhao, Q. Chen, Y. Zhang, L. Zheng, K.C. Chou, Glioma stages prediction based on machine learning algorithm combined with protein-protein interaction networks. Genomics doi: 10.1016/j.ygeno.2019.05.024Get (2019).
- J. Jia, Z. Liu, X. Xiao, B. Liu, K.C. Chou, Identification of protein-protein binding sites by incorporating the physicochemical properties and stationary wavelet transforms into pseudo amino acid composition (iPPBS-PseAAC). J Biomol Struct Dyn (JBSD) 34 (2016) 1946–1961.
- J. Jia, Z. Liu, X. Xiao, B. Liu, K.C. Chou, iSuc-PseOpt: Identifying lysine succinylation sites in proteins by incorporating sequence-coupling effects into pseudo components and optimizing imbalanced training dataset. Anal Biochem 497 (2016) 48–56.
- J. Jia, Z. Liu, X. Xiao, B. Liu, K.C. Chou, pSuc-Lys: Predict lysine succinylation sites in proteins with PseAAC and ensemble random forest approach. Journal of Theoretical Biology 394 (2016) 223–230.
- J. Jia, Z. Liu, X. Xiao, B. Liu, K.C. Chou, iCar-PseCp: identify carbonylation sites in proteins by Monto Carlo sampling and incorporating sequence coupled effects into general PseAAC. Oncotarget 7 (2016) 34558–34570.
- J. Jia, Z. Liu, X. Xiao, B. Liu, K.C. Chou, iPPBS-Opt: A Sequence-Based Ensemble Classifier for Identifying Protein-Protein Binding Sites by Optimizing Imbalanced Training Datasets. Molecules 21 (2016) E95.
- J. Jia, L. Zhang, Z. Liu, X. Xiao, K.C. Chou, pSumo-CD: Predicting sumoylation sites in proteins with covariance discriminant algorithm by incorporating sequence-coupled effects into general PseAAC. Bioinformatics 32 (2016) 3133–3141.
- Z. Liu, X. Xiao, D.J. Yu, J. Jia, W.R. Qiu, K.C. Chou, pRNAm-PC: Predicting N-methyladenosine sites in RNA sequences via physical-chemical properties. Anal Biochem 497 (2016) 60–67.
- X. Xiao, H.X. Ye, Z. Liu, J.H. Jia, K.C. Chou, iROS-gPseKNC: predicting replication origin sites in DNA by incorporating dinucleotide position-specific propensity into general pseudo nucleotide composition. Oncotarget 7 (2016) 34180–34189.
- W.R. Qiu, B.Q. Sun, X. Xiao, Z.C. Xu, J.H. Jia, K.C. Chou, iKcr-PseEns: Identify lysine crotonylation sites in histone proteins with pseudo components and ensemble classifier. Genomics 110 (2018) 239–246.
- J. Jia, X. Li, W. Qiu, X. Xiao, K.C. Chou, iPPI-PseAAC(CGR): Identify protein-protein interactions by incorporating chaos game representation into PseAAC. Journal of Theoretical Biology 460 (2019) 195–203.
- W. Chen, H. Ding, P. Feng, H. Lin, K.C. Chou, iACP: a sequence-based tool for identifying anticancer peptides. Oncotarget 7 (2016) 16895–16909.
- W. Chen, P. Feng, H. Ding, H. Lin, K.C. Chou, Using deformation energy to analyze nucleosome positioning in genomes. Genomics 107 (2016) 69–75.
- W. Chen, H. Tang, J. Ye, H. Lin, K.C. Chou, iRNA-PseU: Identifying RNA pseudouridine sites Molecular Therapy - Nucleic Acids 5 (2016) e332.
- C.J. Zhang, H. Tang, W.C. Li, H. Lin, W. Chen, K.C. Chou, iOri-Human: identify human origin of replication by incorporating dinucleotide physicochemical properties into pseudo nucleotide composition. Oncotarget 7 (2016) 69783–69793.
- W. Chen, P. Feng, H. Yang, H. Ding, H. Lin, K.C. Chou, iRNA-AI: identifying the adenosine to inosine editing sites in RNA sequences. Oncotarget 8 (2017) 4208–4217.
- P. Feng, H. Ding, H. Yang, W. Chen, H. Lin, K.C. Chou, iRNA-PseColl: Identifying the occurrence sites of different RNA modifications by incorporating collective effects of nucleotides into PseKNC. Molecular Therapy - Nucleic Acids 7 (2017) 155–163.
- W. Chen, H. Ding, X. Zhou, H. Lin, K.C. Chou, iRNA(m6A)-PseDNC: Identifying N6-methyladenosine sites using pseudo dinucleotide composition. Analytical Biochemistry 561–562 (2018) 59–65.
- W. Chen, P. Feng, H. Yang, H. Ding, H. Lin, K.C. Chou, iRNA-3typeA: identifying 3-types of modification at RNA’s adenosine sites. Molecular Therapy: Nucleic Acid 11 (2018) 468–474.
- Z.D. Su, Y. Huang, Z.Y. Zhang, Y.W. Zhao, D. Wang, W. Chen, K.C. Chou, H. Lin, iLoc-lncRNA: predict the subcellular location of lncRNAs by incorporating octamer composition into general PseKNC. Bioinformatics 34 (2018) 4196–4204.
- H. Yang, W.R. Qiu, G. Liu, F.B. Guo, W. Chen, K.C. Chou, H. Lin, iRSpot-Pse6NC: Identifying recombination spots in Saccharomyces cerevisiae by incorporating hexamer composition into general PseKNC International Journal of Biological Sciences 14 (2018) 883–891.
- P. Feng, H. Yang, H. Ding, H. Lin, W. Chen, K.C. Chou, iDNA6mA-PseKNC: Identifying DNA N(6)-methyladenosine sites by incorporating nucleotide physicochemical properties into PseKNC. Genomics 111 (2019) 96–102.
- Q.S. Du, S.Q. Wang, N.Z. Xie, Q.Y. Wang, R.B. Huang, K.C. Chou, 2L-PCA: A two-level principal component analyzer for quantitative drug design and its applications. Oncotarget 8 (2017) 70564–70578.
- B. Liu, L. Fang, R. Long, X. Lan, K.C. Chou, iEnhancer-2L: a two-layer predictor for identifying enhancers and their strength by pseudo k-tuple nucleotide composition. Bioinformatics 32 (2016) 362–369.
- B. Liu, R. Long, K.C. Chou, iDHS-EL: Identifying DNase I hypersensi-tivesites by fusing three different modes of pseudo nucleotide composition into an ensemble learning framework. Bioinformatics 32 (2016) 2411–2418.
- B. Liu, S. Wang, R. Long, K.C. Chou, iRSpot-EL: identify recombination spots with an ensemble learning approach. Bioinformatics 33 (2017) 35–41.
- B. Liu, F. Yang, K.C. Chou, 2L-piRNA: A two-layer ensemble classifier for identifying piwi-interacting RNAs and their function. Molecular Therapy - Nucleic Acids 7 (2017) 267–277.
- B. Liu, K. Li, D.S. Huang, K.C. Chou, iEnhancer-EL: Identifying enhancers and their strength with ensemble learning approach. Bioinformatics 34 (2018) 3835–3842.
- B. Liu, F. Weng, D.S. Huang, K.C. Chou, iRO-3wPseKNC: Identify DNA replication origins by three-window-based PseKNC. Bioinformatics 34 (2018) 3086–3093.
- B. Liu, F. Yang, D.S. Huang, K.C. Chou, iPromoter-2L: a two-layer predictor for identifying promoters and their types by multi-window-based PseKNC. Bioinformatics 34 (2018) 33–40.
- W.R. Qiu, B.Q. Sun, X. Xiao, Z.C. Xu, K.C. Chou, iHyd-PseCp: Identify hydroxyproline and hydroxylysine in proteins by incorporating sequence-coupled effects into general PseAAC. Oncotarget 7 (2016) 44310–44321.
- W.R. Qiu, B.Q. Sun, X. Xiao, Z.C. Xu, K.C. Chou, iPTM-mLys: identifying multiple lysine PTM sites and their different types. Bioinformatics 32 (2016) 3116–3123.
- W.R. Qiu, X. Xiao, Z.C. Xu, K.C. Chou, iPhos-PseEn: identifying phosphorylation sites in proteins by fusing different pseudo components into an ensemble classifier. Oncotarget 7 (2016) 51270–51283.
- W.R. Qiu, S.Y. Jiang, B.Q. Sun, X. Xiao, X. Cheng, K.C. Chou, iRNA-2methyl: identify RNA 2’-O-methylation sites by incorporating sequence-coupled effects into general PseKNC and ensemble classifier. Medicinal Chemistry 13 (2017) 734–743.
- W.R. Qiu, S.Y. Jiang, Z.C. Xu, X. Xiao, K.C. Chou, iRNAm5C-PseDNC: identifying RNA 5-methylcytosine sites by incorporating physical-chemical properties into pseudo dinucleotide composition. Oncotarget 8 (2017) 41178–41188.
- W.R. Qiu, B.Q. Sun, X. Xiao, D. Xu, K.C. Chou, iPhos-PseEvo: Identifying human phosphorylated proteins by incorporating evolutionary information into general PseAAC via grey system theory. Molecular Informatics 36 (2017) UNSP 1600010.
- X. Zhai, M. Chen, W. Lu, Accelerated search for perovskite materials with higher Curie temperature based on the machine learning methods. Computational Materials Science 151 (2018) 41–48.
- K.C. Chou, Some remarks on protein attribute prediction and pseudo amino acid composition (50th Anniversary Year Review, 5-steps rule). Journal of Theoretical Biology 273 (2011) 236–247.
- K.C. Chou, Impacts of pseudo amino acid components and 5-steps rule to proteomics and proteome analysis. Current Topics in Medicinak Chemistry (CTMC) (Special Issue ed. G.P Zhou) https://doi.org/10.2174/1568026619666191018100141 or http://www.eurekaselect.com/175823/article (2019).
- K.C. Chou, Prediction of protein cellular attributes using pseudo amino acid composition. PROTEINS: Structure, Function, and Genetics (Erratum: ibid., 2001, Vol.44, 60) 43 (2001) 246–255.
- K.C. Chou, Using amphiphilic pseudo amino acid composition to predict enzyme subfamily classes. Bioinformatics 21 (2005) 10–19.
- K.C. Chou, Pseudo amino acid composition and its applications in bioinformatics, proteomics and system biology. Current Proteomics 6 (2009) 262–274.
- Y.S. Ding, T.L. Zhang, Using Chou’s pseudo amino acid composition to predict subcellular localization of apoptosis proteins: an approach with immune genetic algorithm-based ensemble classifier. Pattern Recognition Letters 29 (2008) 1887–1892.
- Y. Fang, Y. Guo, Y. Feng, M. Li, Predicting DNA-binding proteins: approached from Chou’s pseudo amino acid composition and other specific sequence features. Amino Acids 34 (2008) 103–109.
- X. Jiang, R. Wei, T.L. Zhang, Q. Gu, Using the concept of Chou’s pseudo amino acid composition to predict apoptosis proteins subcellular location: an approach by approximate entropy. Protein & Peptide Letters 15 (2008) 392–396.
- X. Jiang, R. Wei, Y. Zhao, T. Zhang, Using Chou’s pseudo amino acid composition based on approximate entropy and an ensemble of AdaBoost classifiers to predict protein subnuclear location. Amino Acids 34 (2008) 669–675.
- F.M. Li, Q.Z. Li, Predicting protein subcellular location using Chou’s pseudo amino acid composition and improved hybrid approach. Protein & Peptide Letters 15 (2008) 612–616.
- H. Lin, The modified Mahalanobis discriminant for predicting outer membrane proteins by using Chou’s pseudo amino acid composition. Journal of Theoretical Biology 252 (2008) 350–356.
- H. Lin, H. Ding, F.B. Feng-Biao Guo, A.Y. Zhang, J. Huang, Predicting subcellular localization of mycobacterial proteins by using Chou’s pseudo amino acid composition. Protein & Peptide Letters 15 (2008) 739–744.
- L. Nanni, A. Lumini, Genetic programming for creating Chou’s pseudo amino acid based features for submitochondria localization. Amino Acids 34 (2008) 653–660.
- G.Y. Zhang, H.C. Li, J.Q. Gao, B.S. Fang, Predicting lipase types by improved Chou’s pseudo amino acid composition. Protein & Peptide Letters 15 (2008) 1132–1137.
- S.W. Zhang, W. Chen, F. Yang, Q. Pan, Using Chou’s pseudo amino acid composition to predict protein quaternary structure: a sequence-segmented PseAAC approach. Amino Acids 35 (2008) 591–598.
- S.W. Zhang, Y.L. Zhang, H.F. Yang, C.H. Zhao, Q. Pan, Using the concept of Chou’s pseudo amino acid composition to predict protein subcellular localization: an approach by incorporating evolutionary information and von Neumann entropies. Amino Acids 34 (2008) 565–572.
- C. Chen, L. Chen, X. Zou, P. Cai, Prediction of protein secondary structure content by using the concept of Chou’s pseudo amino acid composition and support vector machine. Protein & Peptide Letters 16 (2009) 27–31.
- D.N. Georgiou, T.E. Karakasidis, J.J. Nieto, A. Torres, Use of fuzzy clustering technique and matrices to classify amino acids and its impact to Chou’s pseudo amino acid composition. Journal of Theoretical Biology 257 (2009) 17–26.
- Z.C. Li, X.B. Zhou, Z. Dai, X.Y. Zou, Prediction of protein structural classes by Chou’s pseudo amino acid composition: approached using continuous wavelet transform and principal component analysis. Amino Acids 37 (2009) 415–425.
- H. Lin, H. Wang, H. Ding, Y.L. Chen, Q.Z. Li, Prediction of Subcellular Localization of Apoptosis Protein Using Chou’s Pseudo Amino Acid Composition. Acta Biotheoretica 57 (2009) 321–330.
- J.D. Qiu, J.H. Huang, R.P. Liang, X.Q. Lu, Prediction of G-protein-coupled receptor classes based on the concept of Chou’s pseudo amino acid composition: an approach from discrete wavelet transform. Analytical Biochemistry 390 (2009) 68–73.
- Y.H. Zeng, Y.Z. Guo, R.Q. Xiao, L. Yang, L.Z. Yu, M.L. Li, Using the augmented Chou’s pseudo amino acid composition for predicting protein submitochondria locations based on auto covariance approach. Journal of Theoretical Biology 259 (2009) 366–372.
- M. Esmaeili, H. Mohabatkar, S. Mohsenzadeh, Using the concept of Chou’s pseudo amino acid composition for risk type prediction of human papillomaviruses. Journal of Theoretical Biology 263 (2010) 203–209.
- Q. Gu, Y.S. Ding, T.L. Zhang, Prediction of G-Protein-Coupled Receptor Classes in Low Homology Using Chou’s Pseudo Amino Acid Composition with Approximate Entropy and Hydrophobicity Patterns. Protein & Peptide Letters 17 (2010) 559–567.
- H. Mohabatkar, Prediction of cyclin proteins using Chou’s pseudo amino acid composition. Protein & Peptide Letters 17 (2010) 1207–1214.
- J.D. Qiu, J.H. Huang, S.P. Shi, R.P. Liang, Using the concept of Chou’s pseudo amino acid composition to predict enzyme family classes: an approach with support vector machine based on discrete wavelet transform. Protein & Peptide Letters 17 (2010) 715–722.
- S.S. Sahu, G. Panda, A novel feature representation method based on Chou’s pseudo amino acid composition for protein structural class prediction. Computational Biology and Chemistry 34 (2010) 320–327.
- L. Yu, Y. Guo, Y. Li, G. Li, M. Li, J. Luo, W. Xiong, W. Qin, SecretP: Identifying bacterial secreted proteins by fusing new features into Chou’s pseudo amino acid composition. Journal of Theoretical Biology 267 (2010) 1–6.
- J. Guo, N. Rao, G. Liu, Y. Yang, G. Wang, Predicting protein folding rates using the concept of Chou’s pseudo amino acid composition. Journal of Computational Chemistry 32 (2011) 1612–1617.
- J. Lin, Y. Wang, Using a novel AdaBoost algorithm and Chou’s pseudo amino acid composition for predicting protein subcellular localization. Protein & Peptide Letters 18 (2011) 1219–1225.
- J. Lin, Y. Wang, X. Xu, A novel ensemble and composite approach for classifying proteins based on Chou’s pseudo amino acid composition. African Journal of Biotechnology 10 (2011) 16963–16968.
- H. Mohabatkar, M. Mohammad Beigi, A. Esmaeili, Prediction of GABA(A) receptor proteins using the concept of Chou’s pseudo amino acid composition and support vector machine. Journal of Theoretical Biology 281 (2011) 18–23.
- B.M. Mohammad, M. Behjati, H. Mohabatkar, Prediction of metalloproteinase family based on the concept of Chou’s pseudo amino acid composition using a machine learning approach. Journal of Structural and Functional Genomics 12 (2011) 191–197.
- J.D. Qiu, S.B. Suo, X.Y. Sun, S.P. Shi, R.P. Liang, OligoPred: A web-server for predicting homo-oligomeric proteins by incorporating discrete wavelet transform into Chou’s pseudo amino acid composition. Journal of Molecular Graphics & Modelling 30 (2011) 129–134.
- D. Zou, Z. He, J. He, Y. Xia, Supersecondary structure prediction using Chou’s pseudo amino acid composition. Journal of Computational Chemistry 32 (2011) 271–278.
- J.Z. Cao, W.Q. Liu, H. Gu, Predicting Viral Protein Subcellular Localization with Chou’s Pseudo Amino Acid Composition and Imbalance-Weighted Multi-Label K-Nearest Neighbor Algorithm. Protein and Peptide Letters 19 (2012) 1163–1169.
- C. Chen, Z.B. Shen, X.Y. Zou, Dual-Layer Wavelet SVM for Predicting Protein Structural Class Via the General Form of Chou’s Pseudo Amino Acid Composition. Protein & Peptide Letters 19 (2012) 422–429.
- P. Du, X. Wang, C. Xu, Y. Gao, PseAAC-Builder: A cross-platform stand-alone program for generating various special Chou’s pseudo amino acid compositions. Analytical Biochemistry 425 (2012) 117–119.
- G.L. Fan, Q.Z. Li, Predict mycobacterial proteins subcellular locations by incorporating pseudo-average chemical shift into the general form of Chou’s pseudo amino acid composition. Journal of Theoretical Biology 304 (2012) 88–95.
- G.L. Fan, Q.Z. Li, Predicting protein submitochondria locations by combining different descriptors into the general form of Chou’s pseudo amino acid composition. Amino Acids 43 (2012) 545–555.
- M. Hayat, A. Khan, Discriminating Outer Membrane Proteins with Fuzzy K-Nearest Neighbor Algorithms Based on the General Form of Chou’s PseAAC. Protein & Peptide Letters 19 (2012) 411–421.
- L.Q. Li, Y. Zhang, L.Y. Zou, Y. Zhou, X.Q. Zheng, Prediction of Protein Subcellular Multi-Localization Based on the General form of Chou’s Pseudo Amino Acid Composition. Protein & Peptide Letters 19 (2012) 375–387.
- B. Liao, Q. Xiang, D. Li, Incorporating Secondary Features into the General form of Chou’s PseAAC for Predicting Protein Structural Class. Protein & Peptide Letters 19 (2012) 1133–1138.
- L. Liu, X.Z. Hu, X.X. Liu, Y. Wang, S.B. Li, Predicting Protein Fold Types by the General Form of Chou’s Pseudo Amino Acid Composition: Approached from Optimal Feature Extractions. Protein & Peptide Letters 19 (2012) 439–449.
- S. Mei, Multi-kernel transfer learning based on Chou’s PseAAC formulation for protein submitochondria localization. Journal of Theoretical Biology 293 (2012) 121–130.
- S. Mei, Predicting plant protein subcellular multi-localization by Chou’s PseAAC formulation based multi-label homolog knowledge transfer learning. Journal of Theoretical Biology 310 (2012) 80–87.
- L. Nanni, S. Brahnam, A. Lumini, Wavelet images and Chou’s pseudo amino acid composition for protein classification. Amino Acids 43 (2012) 657–65.
- L. Nanni, A. Lumini, D. Gupta, A. Garg, Identifying bacterial virulent proteins by fusing a set of classifiers based on variants of Chou’s pseudo amino acid composition and on evolutionary information. IEEE-ACM Transaction on Computational Biolology and Bioinformatics 9 (2012) 467–475.
- X.H. Niu, X.H. Hu, F. Shi, J.B. Xia, Predicting Protein Solubility by the General Form of Chou’s Pseudo Amino Acid Composition: Approached from Chaos Game Representation and Fractal Dimension. Protein & Peptide Letters 19 (2012) 940–948.
- Y.F. Qin, C.H. Wang, X.Q. Yu, J. Zhu, T.G. Liu, X.Q. Zheng, Predicting Protein Structural Class by Incorporating Patterns of Over- Represented k-mers into the General form of Chou’s PseAAC. Protein & Peptide Letters 19 (2012) 388–397.
- L.Y. Ren, Y.S. Zhang, I. Gutman, Predicting the Classification of Transcription Factors by Incorporating their Binding Site Properties into a Novel Mode of Chou’s Pseudo Amino Acid Composition. Protein & Peptide Letters 19 (2012) 1170–1176.
- X.Y. Sun, S.P. Shi, J.D. Qiu, S.B. Suo, S.Y. Huang, R.P. Liang, Identifying protein quaternary structural attributes by incorporating physicochemical properties into the general form of Chou’s PseAAC via discrete wavelet transform. Molecular BioSystems 8 (2012) 3178–3184.
- X.W. Zhao, Z.Q. Ma, M.H. Yin, Predicting protein-protein interactions by combing various sequence- derived features into the general form of Chou’s Pseudo amino acid composition. Protein & Peptide Letters 19 (2012) 492–500.
- Zia-ur-Rehman, A. Khan, Identifying GPCRs and their Types with Chou’s Pseudo Amino Acid Composition: An Approach from Multi-scale Energy Representation and Position Specific Scoring Matrix. Protein & Peptide Letters 19 (2012) 890–903.
- D.S. Cao, Q.S. Xu, Y.Z. Liang, propy: a tool to generate various modes of Chou’s PseAAC. Bioinformatics 29 (2013) 960–962.
- T.H. Chang, L.C. Wu, T.Y. Lee, S.P. Chen, H.D. Huang, J.T. Horng, EuLoc: a web-server for accurately predict protein subcellular localization in eukaryotes by incorporating various features of sequence segments into the general form of Chou’s PseAAC. Journal of Computer-Aided Molecular Design 27 (2013) 91–103.
- Y.K. Chen, K.B. Li, Predicting membrane protein types by incorporating protein topology, domains, signal peptides, and physicochemical properties into the general form of Chou’s pseudo amino acid composition. Journal of Theoretical Biology 318 (2013) 1–12.
- G.-L. Fan, Q.-Z. Li, Y.-C. Zuo, Predicting acidic and alkaline enzymes by incorporating the average chemical shift and gene ontology informations into the general form of Chou’s PseAAC. Pocess Biochemistry 48 (2013) 1048–1053.
- G.L. Fan, Q.Z. Li, Discriminating bioluminescent proteins by incorporating average chemical shift and evolutionary information into the general form of Chou’s pseudo amino acid composition. Journal of Theoretical Biology 334 (2013) 45–51.
- D.N. Georgiou, T.E. Karakasidis, A.C. Megaritis, A short survey on genetic sequences, Chou’s pseudo amino acid composition and its combination with fuzzy set theory. The Open Bioinformatics Journal 7 (2013) 41–48.
- M.K. Gupta, R. Niyogi, M. Misra, An alignment-free method to find similarity among protein sequences via the general form of Chou’s pseudo amino acid composition. SAR QSAR Environ Res 24 (2013) 597–609.
- C. Huang, J. Yuan, Using radial basis function on the general form of Chou’s pseudo amino acid composition and PSSM to predict subcellular locations of proteins with both single and multiple sites. Biosystems 113 (2013) 50–57.
- C. Huang, J.Q. Yuan, A multilabel model based on Chou’s pseudo amino acid composition for identifying membrane proteins with both single and multiple functional types. J Membr Biol 246 (2013) 327–34.
- C. Huang, J.Q. Yuan, Predicting protein subchloroplast locations with both single and multiple sites via three different modes of Chou’s pseudo amino acid compositions. Journal of Theoretical Biology 335 (2013) 205–12.
- M. Khosravian, F.K. Faramarzi, M.M. Beigi, M. Behbahani, H. Mohabatkar, Predicting Antibacterial Peptides by the Concept of Chou’s Pseudo amino Acid Composition and Machine Learning Methods. Protein & Peptide Letters 20 (2013) 180–186.
- H. Lin, C. Ding, L.-F. Yuan, W. Chen, H. Ding, Z.-Q. Li, F.-B. Guo, J. Huang, N.-N. Rao, Predicting subchloroplast locations of proteins based on the general form of Chou’s pseudo amino acid composition: Approached from optimal tripeptide composition. International Journal of Biomethmatics 6 (2013) 1350003.
- B. Liu, X. Wang, Q. Zou, Q. Dong, Q. Chen, Protein remote homology detection by combining Chou’s pseudo amino acid composition and profile-based protein representation. Molecular Informatics 32 (2013) 775–782.
- H. Mohabatkar, M.M. Beigi, K. Abdolahi, S. Mohsenzadeh, Prediction of Allergenic Proteins by Means of the Concept of Chou’s Pseudo Amino Acid Composition and a Machine Learning Approach. Medicinal Chemistry 9 (2013) 133–137.
- E. Pacharawongsakda, T. Theeramunkong, Predict Subcellular Locations of Singleplex and Multiplex Proteins by Semi-Supervised Learning and Dimension-Reducing General Mode of Chou’s PseAAC. IEEE Transactions on Nanobioscience 12 (2013) 311–320.
- Y.F. Qin, L. Zheng, J. Huang, Locating apoptosis proteins by incorporating the signal peptide cleavage sites into the general form of Chou’s Pseudo amino acid composition. International Journal of Quantum Chemistry 113 (2013) 1660–1667.
- A.N. Sarangi, M. Lohani, R. Aggarwal, Prediction of Essential Proteins in Prokaryotes by Incorporating Various Physico-chemical Features into the General form of Chou’s Pseudo Amino Acid Composition. Protein Pept Lett 20 (2013) 781–95.
- S. Wan, M.W. Mak, S.Y. Kung, GOASVM: A subcellular location predictor by incorporating term-frequency gene ontology into the general form of Chou’s pseudo amino acid composition. Journal of Theoretical Biology 323 (2013) 40–48.
- X. Wang, G.Z. Li, W.C. Lu, Virus-ECC-mPLoc: a multi-label predictor for predicting the subcellular localization of virus proteins with both single and multiple sites based on a general form of Chou’s pseudo amino acid composition. Protein & Peptide Letters 20 (2013) 309–317.
- N. Xiaohui, L. Nana, X. Jingbo, C. Dingyan, P. Yuehua, X. Yang, W. Weiquan, W. Dongming, W. Zengzhen, Using the concept of Chou’s pseudo amino acid composition to predict protein solubility: An approach with entropies in information theory. Journal of Theoretical Biology 332 (2013) 211–217.
- H.L. Xie, L. Fu, X.D. Nie, Using ensemble SVM to identify human GPCRs N-linked glycosylation sites based on the general form of Chou’s PseAAC. Protein Eng Des Sel 26 (2013) 735–742.
- P. Du, S. Gu, Y. Jiao, PseAAC-General: Fast building various modes of general form of Chou’s pseudo amino acid composition for large-scale protein datasets. International Journal of Molecular Sciences 15 (2014) 3495–3506.
- Z. Hajisharifi, M. Piryaiee, M. Mohammad Beigi, M. Behbahani, H. Mohabatkar, Predicting anticancer peptides with Chou’s pseudo amino acid composition and investigating their mutagenicity via Ames test. Journal of Theoretical Biology 341 (2014) 34–40.
- G.S. Han, Z.G. Yu, V. Anh, A two-stage SVM method to predict membrane protein types by incorporating amino acid classifications and physicochemical properties into a general form of Chou’s PseAAC. J Theor Biol 344 (2014) 31–9.
- C. Jia, X. Lin, Z. Wang, Prediction of Protein S-Nitrosylation Sites Based on Adapted Normal Distribution Bi-Profile Bayes and Chou’s Pseudo Amino Acid Composition. Int J Mol Sci 15 (2014) 10410–23.
- L. Kong, L. Zhang, J. Lv, Accurate prediction of protein structural classes by incorporating predicted secondary structure information into the general form of Chou’s pseudo amino acid composition. J Theor Biol 344 (2014) 12–18.
- L. Li, S. Yu, W. Xiao, Y. Li, M. Li, L. Huang, X. Zheng, S. Zhou, H. Yang, Prediction of bacterial protein subcellular localization by incorporating various features into Chou’s PseAAC and a backward feature selection approach. Biochimie 104 (2014) 100–7.
- L. Nanni, S. Brahnam, A. Lumini, Prediction of protein structure classes by incorporating different protein descriptors into general Chou’s pseudo amino acid composition. J Theor Biol 360 (2014) 109–116.
- J. Zhang, P. Sun, X. Zhao, Z. Ma, PECM: Prediction of extracellular matrix proteins using the concept of Chou’s pseudo amino acid composition. Journal of Theoretical Biology 363 (2014) 412–418.
- J. Zhang, X. Zhao, P. Sun, Z. Ma, PSNO: Predicting Cysteine S-Nitrosylation Sites by Incorporating Various Sequence-Derived Features into the General Form of Chou’s PseAAC. Int J Mol Sci 15 (2014) 11204–19.
- L. Zhang, X. Zhao, L. Kong, Predict protein structural class for low-similarity sequences by evolutionary difference information into the general form of Chou’s pseudo amino acid composition. J Theor Biol 355 (2014) 105–10.
- Y.C. Zuo, Y. Peng, L. Liu, W. Chen, L. Yang, G.L. Fan, Predicting peroxidase subcellular location by hybridizing different descriptors of Chou’s pseudo amino acid patterns. Anal Biochem 458 (2014) 14–9.
- F. Ali, M. Hayat, Classification of membrane protein types using Voting Feature Interval in combination with Chou’s Pseudo Amino Acid Composition. J Theor Biol 384 (2015) 78–83.
- G.L. Fan, X.Y. Zhang, Y.L. Liu, Y. Nang, H. Wang, DSPMP: Discriminating secretory proteins of malaria parasite by hybridizing different descriptors of Chou’s pseudo amino acid patterns. J Comput Chem 36 (2015) 2317–27.
- C. Huang, J.Q. Yuan, Simultaneously Identify Three Different Attributes of Proteins by Fusing their Three Different Modes of Chou’s Pseudo Amino Acid Compositions. Protein Pept Lett 22 (2015) 547–56.
- Z.U. Khan, M. Hayat, M.A. Khan, Discrimination of acidic and alkaline enzyme using Chou’s pseudo amino acid composition in conjunction with probabilistic neural network model. J Theor Biol 365 (2015) 197–203.
- R. Kumar, A. Srivastava, B. Kumari, M. Kumar, Prediction of beta-lactamase and its class by Chou’s pseudo amino acid composition and support vector machine. J Theor Biol 365 (2015) 96–103.
- B. Liu, J. Xu, S. Fan, R. Xu, J. Jiyun Zhou, X. Wang, PseDNA-Pro: DNA-binding protein identification by combining Chou’s PseAAC and physicochemical distance transformation. Molecular Informatics 34 (2015) 8–17
- M. Mandal, A. Mukhopadhyay, U. Maulik, Prediction of protein subcellular localization by incorporating multiobjective PSO-based feature subset selection into the general form of Chou’s PseAAC. Med Biol Eng Comput 53 (2015) 331–44.
- V. Sanchez, A.M. Peinado, J.L. Perez-Cordoba, A.M. Gomez, A new signal characterization and signal-based Chou’s PseAAC representation of protein sequences. J Bioinform Comput Biol 13 (2015) 1550024.
- X. Wang, W. Zhang, Q. Zhang, G.Z. Li, MultiP-SChlo: multi-label protein subchloroplast localization prediction with Chou’s pseudo amino acid composition and a novel multi-label classifier. Bioinformatics 31 (2015) 2639–45.
- Y.S. Jiao, P.F. Du, Prediction of Golgi-resident protein types using general form of Chou’s pseudo amino acid compositions: Approaches with minimal redundancy maximal relevance feature selection. J Theor Biol 402 (2016) 38–44.
- M. Kabir, M. Hayat, iRSpot-GAEnsC: identifing recombination spots via ensemble classifier and extending the concept of Chou’s PseAAC to formulate DNA samples. Molecular Genetics and Genomics 291 (2016) 285–96.
- M. Tahir, M. Hayat, iNuc-STNC: a sequence-based predictor for identification of nucleosome positioning in genomes by extending the concept of SAAC and Chou’s PseAAC. Mol Biosyst 12 (2016) 2587–93.
- H. Tang, W. Chen, H. Lin, Identification of immunoglobulins using Chou’s pseudo amino acid composition with feature selection technique. Mol Biosyst 12 (2016) 1269–1275.
- H.L. Zou, X. Xiao, Predicting the Functional Types of Singleplex and Multiplex Eukaryotic Membrane Proteins via Different Models of Chou’s Pseudo Amino Acid Compositions. J Membr Biol 249 (2016) 23–9.
- H. Huo, T. Li, S. Wang, Y. Lv, Y. Zuo, L. Yang, Prediction of presynaptic and postsynaptic neurotoxins by combining various Chou’s pseudo components. Sci Rep 7 (2017) 5827.
- Z. Ju, J.J. He, Prediction of lysine propionylation sites using biased SVM and incorporating four different sequence features into Chou’s PseAAC. J Mol Graph Model 76 (2017) 356–363.
- M. Rahimi, M.R. Bakhtiarizadeh, A. Mohammadi-Sangcheshmeh, OOgenesis_Pred: A sequence-based method for predicting oogenesis proteins by six different modes of Chou’s pseudo amino acid composition. J Theor Biol 414 (2017) 128–136.
- P. Tripathi, P.N. Pandey, A novel alignment-free method to classify protein folding types by combining spectral graph clustering with Chou’s pseudo amino acid composition. J Theor Biol 424 (2017) 49–54.
- B. Yu, S. Li, W.Y. Qiu, C. Chen, R.X. Chen, L. Wang, M.H. Wang, Y. Zhang, Accurate prediction of subcellular location of apoptosis proteins combining Chou’s PseAAC and PsePSSM based on wavelet denoising. Oncotarget 8 (2017) 107640–107665.
- B. Yu, L. Lou, S. Li, Y. Zhang, W. Qiu, X. Wu, M. Wang, B. Tian, Prediction of protein structural class for low-similarity sequences using Chou’s pseudo amino acid composition and wavelet denoising. J Mol Graph Model 76 (2017) 260–273.
- J. Ahmad, M. Hayat, MFSC: Multi-voting based Feature Selection for Classification of Golgi Proteins by Adopting the General form of Chou’s PseAAC components. J Theor Biol 463 (2018) 99–109.
- S. Akbar, M. Hayat, iMethyl-STTNC: Identification of N(6)-methyladenosine sites by extending the Idea of SAAC into Chou’s PseAAC to formulate RNA sequences. J Theor Biol 455 (2018) 205–211.
- M.A. Al Maruf, S. Shatabda, iRSpot-SF: Prediction of recombination hotspots by incorporating sequence based features into Chou’s Pseudo components. Genomics doi:10.1016/j.ygeno.2018.06.003 (2018).
- M. Arif, M. Hayat, Z. Jan, iMem-2LSAAC: A two-level model for discrimination of membrane proteins and their types by extending the notion of SAAC into Chou’s pseudo amino acid composition. J Theor Biol 442 (2018) 11–21.
- E. Contreras-Torres, Predicting structural classes of proteins by incorporating their global and local physicochemical and conformational properties into general Chou’s PseAAC. J Theor Biol 454 (2018) 139–145.
- X. Cui, Z. Yu, B. Yu, M. Wang, B. Tian, Q. Ma, UbiSitePred: A novel method for improving the accuracy of ubiquitination sites prediction by using LASSO to select the optimal Chou’s pseudo components. Chemometrics and Intelligent Laboratory Systems (CHEMOLAB) doi:10.1016/j.chemolab.2018.11.012 (2018).
- X. Fu, W. Zhu, B. Liso, L. Cai, L. Peng, J. Yang, Improved DNA-binding protein identification by incorporating evolutionary information into the Chou’s PseAAC. IEEE Access 20 (2018) https://doi.org/10.1109/ACCESS.2018.2876656.
- F. Javed, M. Hayat, Predicting subcellular localizations of multi-label proteins by incorporating the sequence features into Chou’s PseAAC. Genomics https://doi.org/10.1016/j.ygeno.2018.09.004 (2018).
- J. Mei, J. Zhao, Prediction of HIV-1 and HIV-2 proteins by using Chou’s pseudo amino acid compositions and different classifiers. Sci Rep 8 (2018) 2359.
- M. Mousavizadegan, H. Mohabatkar, Computational prediction of antifungal peptides via Chou’s PseAAC and SVM. J Bioinform Comput Biol (2018) 1850016.
- W. Qiu, S. Li, X. Cui, Z. Yu, M. Wang, J. Du, Y. Peng, B. Yu, Predicting protein submitochondrial locations by incorporating the pseudo-position specific scoring matrix into the general Chou’s pseudo-amino acid composition. J Theor Biol 450 (2018) 86–103.
- L. Zhang, L. Kong, iRSpot-ADPM: Identify recombination spots by incorporating the associated dinucleotide product model into Chou’s pseudo components. J Theor Biol 441 (2018) 1–8.
- S. Zhang, Y. Liang, Predicting apoptosis protein subcellular localization by integrating auto-cross correlation and PSSM into Chou’s PseAAC. J Theor Biol 457 (2018) 163–169.
- S. Zhang, K. Yang, Y. Lei, K. Song, iRSpot-DTS: Predict recombination spots by incorporating the dinucleotide-based spare-cross covariance information into Chou’s pseudo components. Genomics 11 (2018) 457–464.
- W. Zhao, L. Wang, T.X. Zhang, Z.N. Zhao, P.F. Du, A brief review on software tools in generating Chou’s pseudo-factor representations for all types of biological sequences. Protein Pept Lett 25 (2018) 822–829.
- J. Ahmad, M. Hayat, MFSC: Multi-voting based feature selection for classification of Golgi proteins by adopting the general form of Chou’s PseAAC components. J Theor Biol 463 (2019) 99–109.
- M.A. Al Maruf, S. Shatabda, iRSpot-SF: Prediction of recombination hotspots by incorporating sequence based features into Chou’s Pseudo components. Genomics 111 (2019) 966–972.
- A.H. Butt, N. Rasool, Y.D. Khan, Prediction of antioxidant proteins by incorporating statistical moments based features into Chou’s PseAAC. Journal of Theoretical Biology 473 (2019) 1–8.
- F. Javed, M. Hayat, Predicting subcellular localization of multi-label proteins by incorporating the sequence features into Chou’s PseAAC. Genomics 111 (2019) 1325–1332.
- Y. Pan, S. Wang, Q. Zhang, Q. Lu, D. Su, Y. Zuo, L. Yang, Analysis and prediction of animal toxins by various Chou’s pseudo components and reduced amino acid compositions. J Theor Biol 462 (2019) 221–229.
- M. Tahir, M. Hayat, S.A. Khan, iNuc-ext-PseTNC: an efficient ensemble model for identification of nucleosome positioning by extending the concept of Chou’s PseAAC to pseudo-tri-nucleotide composition. Mol Genet Genomics 294 (2019) 199–210.
- M. Tahir, H. Tayara, K.T. Chong, iRNA-PseKNC(2methyl): Identify RNA 2’-O-methylation sites by convolution neural network and Chou’s pseudo components. J Theor Biol 465 (2019) 1–6.
- B. Tian, X. Wu, C. Chen, W. Qiu, Q. Ma, B. Yu, Predicting protein-protein interactions by fusing various Chou’s pseudo components and using wavelet denoising approach. J Theor Biol 462 (2019) 329–346.
- L. Zhang, L. Kong, iRSpot-PDI: Identification of recombination spots by incorporating dinucleotide property diversity information into Chou’s pseudo components. Genomics 111 (2019) 457–464.
- S. Zhang, K. Yang, Y. Lei, K. Song, iRSpot-DTS: Predict recombination spots by incorporating the dinucleotide-based spare-cross covariance information into Chou’s pseudo components. Genomics 111 (2019) 1760–1770.
- K.C. Chou, Two kinds of metrics for computational biology. Genomics https://www.sciencedirect.com/science/article/pii/S0888754319304604?via%3Dihub (2019).
- K.C. Chou, Proposing pseudo amino acid components is an important milestone for proteome and genome analyses. International Journal for Peptide Research and Therapeutics (IJPRT) https://doi.org/10.1007/s10989-019-09910-7 or https://link.springer.com/article/10.1007%2Fs10989-019-09910-7 (2019).
- K.C. Chou, An insightful recollection for predicting protein subcellular locations in multi-label systems. Genomics http://doi.org/10.1016/j.ygeno.2019.08.008 or https://www.sciencedirect.com/science/article/pii/S0888754319304604?via%3Dihub (2019).
- K.C. Chou, Progresses in predicting post-translational modification. International Journal of Peptide Research and Therapeutics (IJPRT) https:/doi.org/10.1007/s10989-019-09893-5 or https://link.springer.com/article/10.1007%2Fs10989-019-09893-5 (2019).
- K.C. Chou, Recent Progresses in Predicting Protein Subcellular Localization with Artificial Intelligence (AI) Tools Developed Via the 5-Steps Rule. Japanese Journal of Gastroenterology and Hepatology https:/doi.org/www.jjgastrohepto.org or https://www.jjgastrohepto.org/ (2019).
- K.C. Chou, An insightful recollection since the distorted key theory was born about 23 years ago. Genomics https://doi.org/10.1016/j.ygeno.2019.09.001 or https://www.sciencedirect.com/science/article/pii/S0888754319305543?via%3Dihub (2019).
- K.C. Chou, Artificial intelligence (AI) tools constructed via the 5-steps rule for predicting post-translational modifications. Trends in Artificial Inttelengence (TIA) 3 (2019) 60–74.
- K.C. Chou, Distorted Key Theory and Its Implication for Drug Development. Current Genomics http://www.eurekaselect.com/175823/article or http://www.eurekaselect.com/175823/article (2020).
- K. C. Chou, An Insightful 10-year Recollection Since the Emergence of the 5-steps Rule. Current Pharmaceutical Design 2019, 25, 4223–4234.
- K.C. Chou, An insightful recollection since the birth of Gordon Life Science Institute about 17 years ago. Advancement in Scientific and Engineering Research 4 (2019) 31–36.
- K.C. Chou, Gordon Life Science Institute: Its philosophy, achievements, and perspective. Annals of Cancer Therapy and Pharmacology 2 (2019) 001–26.
Article Type
Short Communication
Publication history
Received: February 15, 2023
Accepted: February 16, 2023
Published: February 19, 2023
Citation:
Kuo-Chen C (2023) The Ploc_Bal-Mgpos is a Powerful Artificial Intelligence Tool for Predicting the Subcellular Localization of Gram-Positive Bacterial Proteins According to Their Sequence Information Alone. Clar J Infect Dis Ther 04(01): 280–292.
Kuo-Chen Chou*
Gordon Life Science Institute, Boston, Massachusetts 02478, USA
*Corresponding author
Kuo-Chen Chou,
Gordon Life Science Institute,
Boston,
Massachusetts 02478,
USA;