- 4192 Devonwood Way, Ashburn, Virginia, 20148
- Helpline: +1 (703) 665-3747
Silencing SATB1 Gene Suppressed Expression of mP53, S1004A Genes and Increased Expression of β-catenin and E-cadherin Genes in Murine Mammary Cancer Cells in the Presence and Absence of Tocotrienol-Rich Fraction (TRF)
- Home
- Back to Journal
- Article Details
Abstract
Tocotrienol-Rich Fraction (TRF), which is vitamin E from palm oil, has been reported to have anticancer effects. Previous studies have shown that TRF supplementation downregulates the expression of special AT-rich sequence-binding protein 1 (SATB1) gene in murine Breast Cancer cells (BC). SATB1 is a unique AT-rich binding protein, which coordinates expression of a number of genes responsible for promoting metastasis in cancer cells. In this study, the SATB1 gene was silenced in the 4T1 murine mammary cancer cell line using the small-interfering RNA (siRNA) approach. Treating the Wild-Type (WT) and SATB1-silenced 4T1 (SATB1–4T1) cells resulted in marked reduction of proliferation in the presence and absence of TRF. The expression of various tumour marker genes (mP53, S1004A) was reduced (p<0.05) in the 4T1 cells that were cultured in the presence or absence of TRF. In addition, the expression of some tumour suppressor genes (β-catenin and E-cadherin) was increased (p < 0.05) in the 4T1 cells that were cultured in the presence or absence of TRF. The findings from this study show that SATB1 gene plays a key role in carcinogenesis in this BC cell line.
Keywords: SiRNA, SATB1, breast cancer, siRNA, knockdown gene
Introduction
Breast cancer (BC) is a major health concern as it is the most common cause of cancer-related deaths in women worldwide. Despite recent developments in BC detection and treatment, there are almost 1.38 million women affected by this disease each year [1]. Some of the challenges in developing effective BC therapies include the complexity of this disease and it ability to rapidly metastasise. Gene silencing is a technique that aims to reduce or eliminate production of a protein by disrupting the gene that is responsible for its expression [2]. There are various gene silencing methods currently employed in research. Some are also being evaluated for its potential as therapeutic agents. Nearly all of these gene silencing approaches involve disabling the function of messenger RNA (mRNA) and prevent it from being translated into the corresponding protein. However, these approaches differ in design, molecule used to disrupt mRNA and the manner of mRNA breakdown [3].
New developments in gene therapy have highlighted the use of small interfering RNA (siRNA) technology as a possible tool to treat breast cancer (BC) and other types of cancers [4]. The SiRNAs are small molecules, which are synthetically made double-stranded RNA (dsRNA) molecules designed to pair with specific mRNA [5]. Binding of SiRNA to its target mRNAs will attract proteins that can result in degradation of the mRNA; thus, preventing the target mRNA from being translated into protein. For siRNA to be a useful therapeutic tool, siRNA-mediated transcriptional silencing has to be efficient, specific and can inhibit tumour growth [6]. Although the SiRNA approach has some limitations, it nevertheless provides a broad potential as a promising tool that can be used to regulate disease-associated genes; including oncogenes and tumour suppressor genes.
Vitamin E from palm oil is known as Tocotrienol-Rich Fraction (TRF). Recently, we reported that the SATB1 gene was downregulated in tumour tissue isolated from TRF supplemented mice induced with BC and treated with Dendritic Cell (DC) vaccine [10]. That was a novel finding, which showed the possible anti-cancer role of TRF as SATB1 gene is reported to play a critical role in the growth and metastasis of BC. In the present study, we investigated the role of SATB1 expression in 4T1 murine mammary cancer cells using the siRNA approach in the presence and absence of TRF. The objective of this study was to investigate the impact of silencing the SATB1 gene on various genes reported to be related to cancer and/or metastasis in the presence and absence of TRF. The results of this study may be useful in developing future cancer therapy using SiRNA-based treatment.
Materials and Methods
Cell Line
4T1 cells, which is a highly metastatic murine mammary cancer cell line was purchased from the American Type Culture Collection (ATCC, USA) and cultured as recommended by the ATCC. The cells were cultured at 37oC in a humidified 5% CO2 incubator (Heraeus, Germany) as described previously [7, 9 & 10].
Silencing of SATB1 Gene
Four siRNA probes (Table 1) specific for murine 4T1 genes were designed and used to transfect 4T1 cells to silence the expression of the SATB1 gene. Transfection was performed using the Hi-Perfect Transfection kit (QIAGEN, Germany). Briefly, the 4T1 cells were seeded at 5 × 104 cells per well in a 24-well plate in 0.5 mL of complete medium (RPMI 1640 supplemented with 10% FBS, 1% glutamine and 1% penicillin/streptomycin). The cells were incubated at 37°C overnight in a humidified 5% CO2 incubator. Three concentrations (20 nM, 30 nM and 40 nM) of each siRNA were evaluated for silencing efficiency, which were freshly prepared in 100 mL of culture medium without FBS before the transfection step. Then, 3 µL Hi-Perfect Transfection reagent was added to the freshly prepared siRNA and the contents were mixed by vortexing. The mixture was incubated for 10 min at room temperature to allow formation of transfection complexes. Following this, the transfection complex was added drop-wise onto the pre-seeded 4T1 cells. The plate was gently swirled to ensure uniform distribution of the transfection complex and the cells were incubated at 37°C overnight in a humidified 5% CO2 for 72 hours. At the end of the culture period, the cells were harvested and RNA was extracted for quantitative PCR (qPCR) analysis to confirm that that the SATB1 gene was silenced.
Table 1: The siRNA premix sequence used for siRNA.
|
No. |
FlexiTube siRNA Premix |
Sequence |
|
|
1 |
Mm_Satb1_5 |
CCCGAAGTACACCATCATCAA |
All siRNA premix lyophilised pellets were diluted with RNase-free water to 625 µL to obtain the concentration of 20 µM |
|
2 |
Mm_Satb1_7 |
AAGGTGGTACAAACATTTCAA |
|
|
3 |
Mm_Satb1_6 |
CAGGAAATGAAGCGTGCTAAA |
|
|
4 |
Mm_Satb1_1 |
CACAGTTGCCCTAAACTAGAA |
Gene Expression Studies
Total RNA was extracted from WT and SATB1-silenced (SATB1-siRNA) 4T1 cells using the Tri-reagent total RNA extraction kit (Molecular Research Centre, USA). The RNA samples were analysed using quantitative real-time PCR (qPCR) to confirm silencing of the SATB1 gene in the 4T1 cells. The QuantiFast SYBR Green RT-PCR master mix (QIAGEN, Germany) used to prepare the RNA samples for qPCR analysis. The qPCR was performed using two-step cycling (Table 2) using a multicolour real-time PCR detection system (IQ5 Hercules, California, USA). The SATB1 and beta-actin genes were purchased from a commercial source (QIAGEN, Germany). The other primers used for qPCR analysis of murine metastatin-1 (Mts-1 S100A4), E-cadherin, B-catenin and mutant p53 (mP53) genes (Table 3) were synthesised using the First Base Sequencing system (Applied Biosystem Inc., USA).
Table 2: Conditions used for the real-time PCR.
|
Step |
Time |
Temperature |
Ramp rate |
|
Reverse transcription |
10 min |
50°C |
|
|
PCR initial activation step |
5 min |
95°C |
Maximal/Fast mode |
|
Two-step cycling |
|
|
|
|
Denaturation |
10sec |
95°C |
Maximal/Fast mode |
|
Combined/annealing/extension |
30 sec |
60°C |
Maximal/Fast mode, perform fluorescence data collection |
|
Number of cycles |
40 |
|
|
|
Last step |
Infinite(∞) |
10°C |
Until take out from cycler. Kept all samples stable |
Table 3: Primers used for the qRT-PCR.
|
Genes (Mouse) |
Reverse /Forward primers |
Reference |
|
Beta-actin |
5’-AACTCGAGTGCCATGGTAACCGTTGAGAC-3’ |
QIAGEN, Germany |
|
5’-AACTCGAGAGACTCCTCAGATGAAGTGTT-3’ |
||
|
S100A4 (mts1) |
5’-AACTCGAGTGCCATGGTAACCGTTGAGAC-3’ |
EL Naaman et. al. 2004 [11] |
|
5’-AACTCGAGAGACTCCTCAGATGAAGTGTT-3’ |
||
|
E-cadherin |
5’-AATGGCGGCAATGCAATCCCAAGA-3’ |
Criswell et.al. 2007 [13] |
|
5’-TGCCACAGACCGATTGTGGAGATA-3’ |
||
|
B-catenin |
5’-TTAAACTCCTGCACCCACCAT-3’ |
Rui Zhang et. al. 2007 [14] |
|
5’-AGGGCAAGGTTTCGAATCAA-3’ |
||
|
p53 |
5’-AAAACCACTTGATGGAGAGTATTTCA-3’ |
Rui Zhang et.al. 2007 [14] |
|
5’-GCTCCCGGAACATCTCGAA-3’ |
||
|
SATB1 |
5’-ACACAGCTCTGCTGCCCAAGCC-3’ |
QIAGEN, Germany |
|
5’-GACCAGCTGAGGACTGATCGG-3’ |
Cell Proliferation
The 4T1 cells were treated with the concentration of the siRNA complex that caused the highest silencing of the SATB1 gene expression in the 4T1 cells. Transfection was performed in 10 mm Upcell Nunc dish (Nunc, Germany). Following incubation of the 4T1 cells with the siRNA complex and Hi-Perfect transfection reagent, the cells were washed with sterile PBS solution and recovered by centrifugation (1500 rpm for 5 min). The cells were seeded at 0.5 x 104 cells/well in complete medium and cultured at 37°C in a humidified 5% CO2 incubator (Heraceus, Germany) overnight. The next day, the culture medium was removed and replaced with a fresh medium, which contained different concentrations (4, 8 or 12 µg/mL) of TRF. Similarly treated WT 4T1 cells were used as controls. The cells were cultured for 72 hours at 37°C in a humidified 5% CO2 incubator (Heraceus, Germany). Following 72 hours, the WT and SATB1-silenced 4T1 cells were washed with RNA-free PBS and RNA was extracted from these cells. The extracted RNA was used for qPCR analysis to check for expression of SATB1 gene and other genes that play a role in carcinogenesis of BC.
Statistical Analysis
Statistical analysis was done using the student T-test and SPSS version 9.12 for most of the data and data represent mean value with + S.E.M of triplicate measurements for qPCR experiments. Results were considered statistically significant when p <0.05.
Results
Optimal conditions for using short interfering RNA approach in 4T1 cells
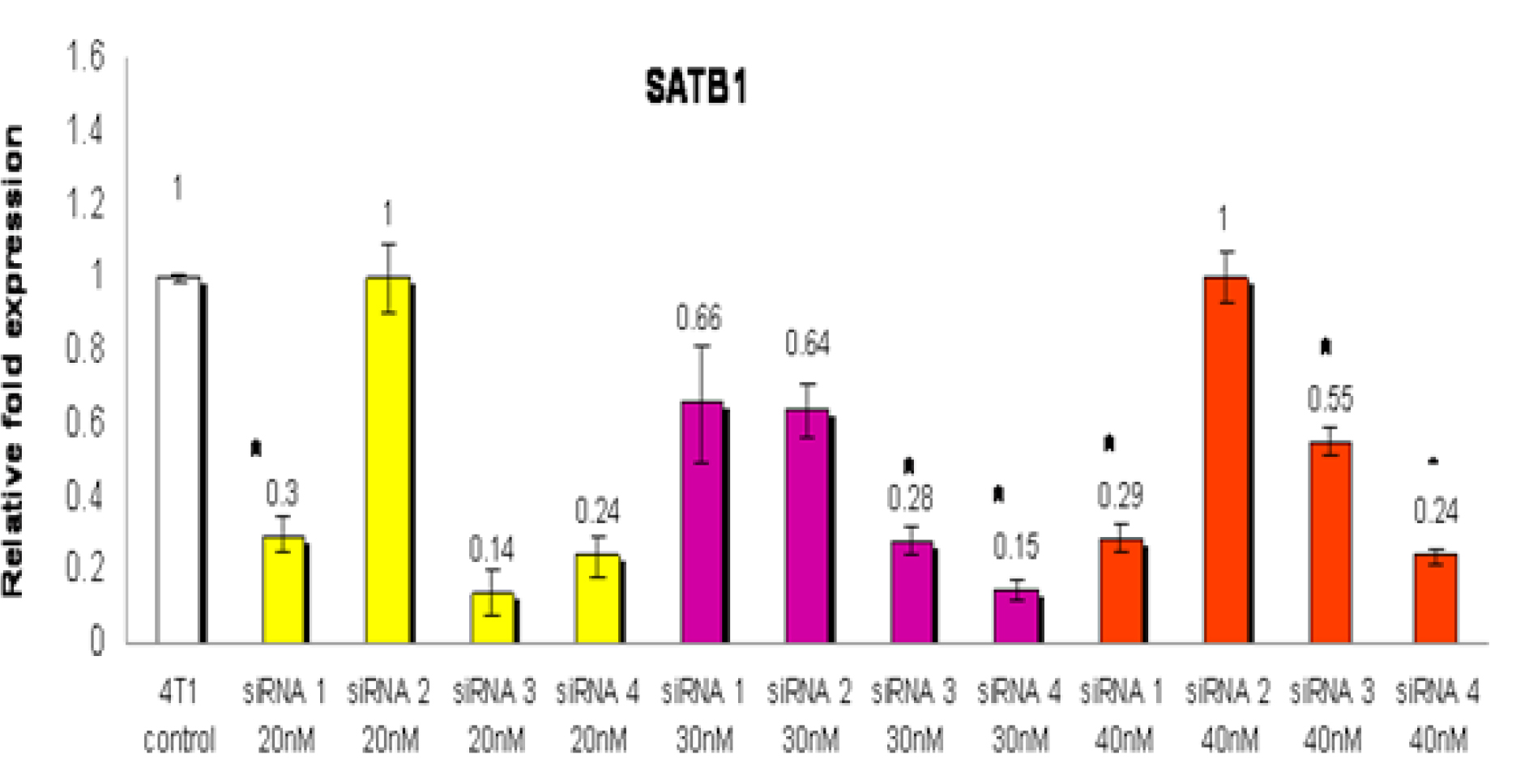
The 4T1 cells exposed to 20 nM siRNA3 showed the most potent inhibition of SATB1 expression (relative fold expression: 0.14) compared to siRNA 1 (relative fold expression: 0.3) and siRNA4 (relative fold expression: 0.24) (Figure 1). However, only the inhibition observed with siRNA1 was found to be statistically significant (p<0.05) when compared with wild-type (control). The relative expression fold when 30 nM siRNA was used were 0.66: 0.64; 0.28 and 0.15 for siRNA1, 2, 3 and 4, respectively (Figure 1). In addition, the inhibition observed with siRNA3 and siRNA4 were found to be statistically significant (p<0.05) when compared with wild-type (control). When 40 nM siRNA was used, the relative expression fold was 0.29: 1.0; 0.55 and 0.24 for siRNA1, 2, 3 and 4, respectively (Figure 1). In addition, the inhibition observed with siRNA1, siRNA3 and siRNA4 were found to be statistically significant (p<0.05) when compared with wild-type (control). From the optimisation step, siRNA 4 at 30nM concentration was found to have the most significant (p<0.05) reduction of the SATB1 gene expression in the 4T1 cells. So, for the next few studies, 30 nM of siRNA 4 was used to silence the expression of the SATB1 gene the 4T1 cells.
Figure 1: The 4T1 cells were treated with four types of siRNA (siRNA 1, siRNA 2, siRNA 3 or siRNA 4) to silence the expression of the SATB1 gene. For each type of siRNA, three different concentrations (20, 30 or 40 nM) of the siRNA were used to identify the siRNA and optimal concentration that gave the highest inhibition of the SATB1 gene expression in the 4T1 cells. RNA from wild-type (WT) 4T1 cells were used as control. Each data point represents mean ± SEM of triplicate samples. [* significantly different from control (p<0.05)]
Viability of SATB1-silenced 4T1 cells treated with TRF
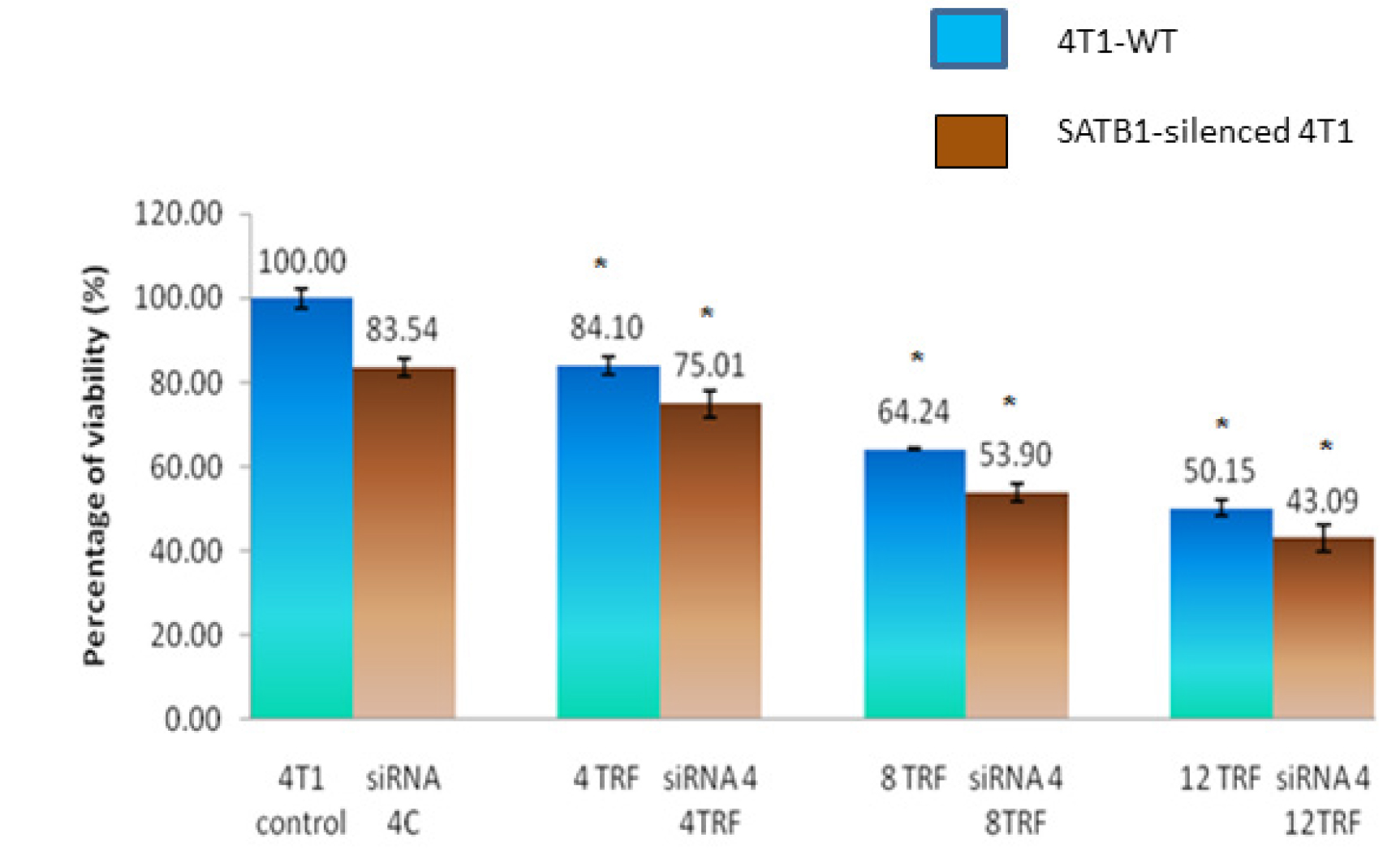
The viability of the SATB1-silenced 4T1 cells was lower (84.1%) when compared to WT 4T1 cells (100%) (Figure 2). In addition, exposure to various concentrations of TRF caused significant reduction (p<0.05) in the viability of the WT and the SATB1-silenced 4T1 (Figure 2). Cell viability reduced when higher concentrations of TRF were used, suggesting that the effects observed were dose-dependent. Overall, the viability of the SATB1-silenced 4T1 cells appears to be lower than the WT 4T1 cells at all concentrations tested.
Figure 2: Wild-type (WT) or SATB1-silenced 4T1 cells (siRNA 4) cells were cultured in the presence or absence of TRF (0, 4, 8 or 12 mg/mL) for 72 hours at 37oC in a humidified 5% CO2 incubator. Cell viability was quantified using the MTT assay. Results are expressed as percentage of cell viability based on untreated controls. Each data point represents mean ± SEM of triplicate samples. [* significantly different from control (p<0.05)]
Expression of SATB1
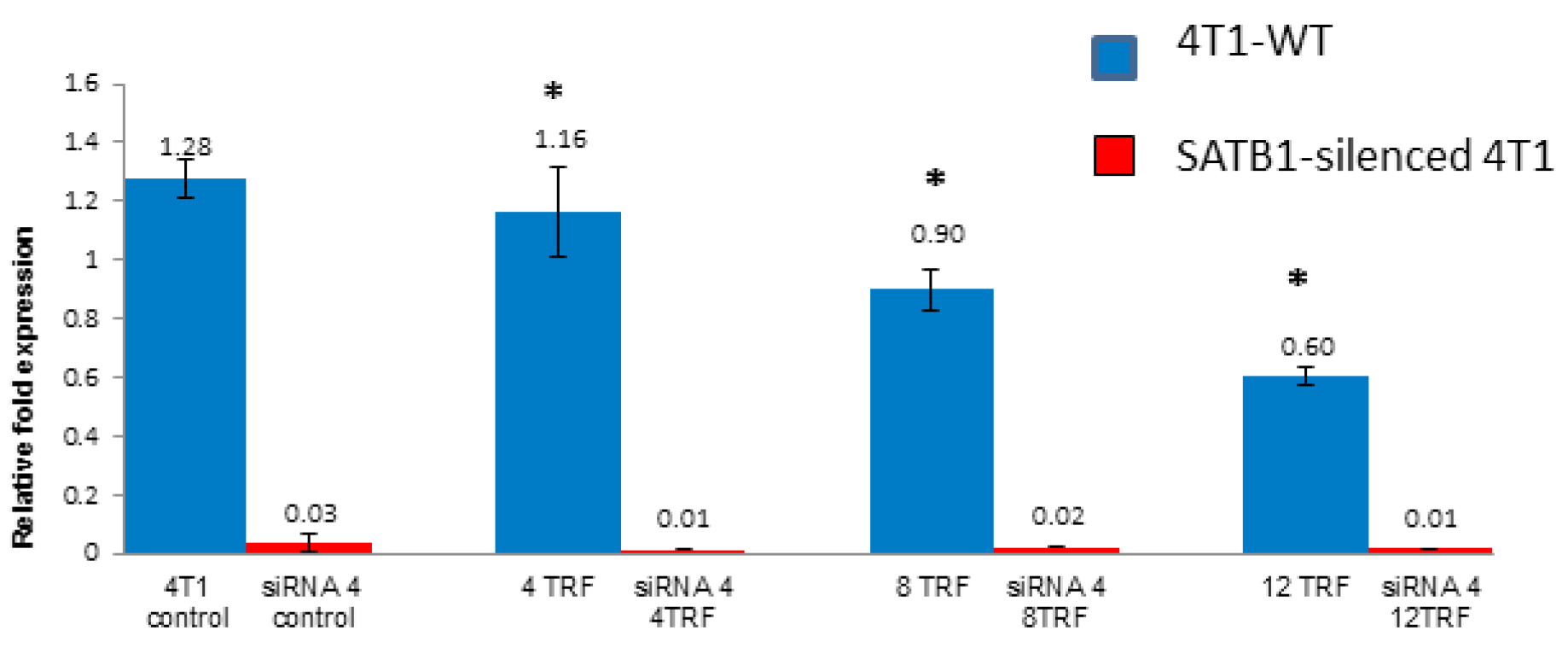
In the siRNA-treated 4T1 cells, there was a marked (p<0.05) down-regulation of the SATB1 gene regardless of exposure to TRF (Figure 3). There were minimal differences observed between each TRF treatments in the SATB1-silenced cells, possibly due to the extremely low expression of the SATB1 gene.
Figure 3: The expression of the SATB1 gene in the wild type (Control) and SATB1-silenced (siRNA 4) 4T1 cells cultured in the presence or absence of TRF (0, 4, 8 or 12 mg/mL) for 72 hours at 37oC in a humidified 5% CO2 incubator was quantified using quantitative PCR. Triplicate preparations were used for each sample. [* significantly different from control (p<0.05)]
Expression of S1004A and mutant P53
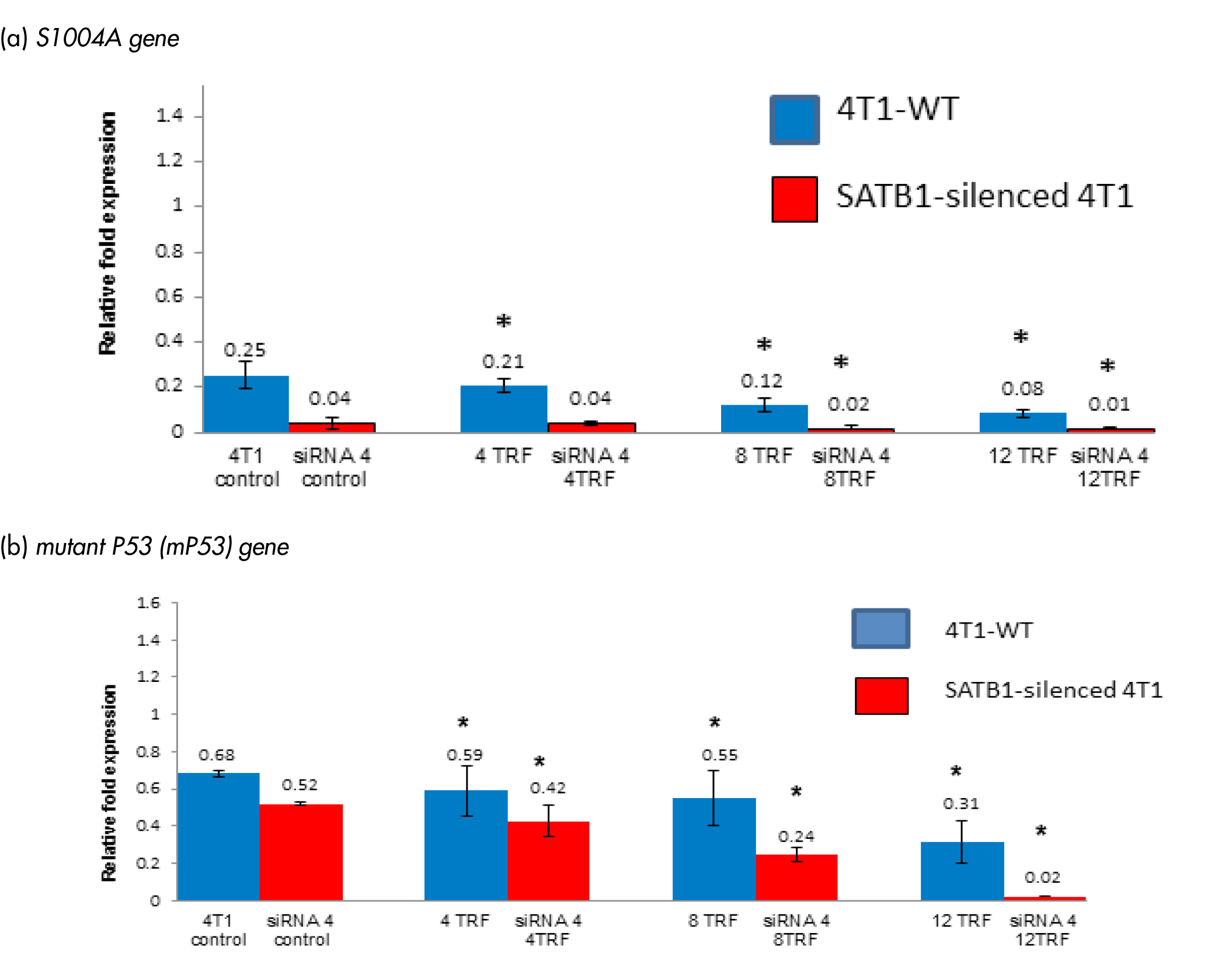
S1004A gene is a metastasis inducer, which is an important regulator of BC progression [16]. This S1004A gene was reported as a SATB1-dependent gene in BC cells [16]. Treating WT 4T1 cells with TRF, down-regulated the expression of the S1004A gene in a dose-dependent manner (Figure 4a). A similar inhibition of the S1004A gene expression was also observed in SATB1-silenced 4T1. Mutant p53 (mP53) is reported to contribute to various stages of tumour progression as well as cause increased resistance to anti-cancer treatments [17]. The expression of the mP53 gene was reduced in WT treated with TRF and the SATB1-silenced 4T1 cells (Figure 4b). Highest level of suppression of the mP53 gene was observed when 12 µg/mL (relative fold expression: 0.31 fold) was used. This was followed by 8 µg/mL (relative fold expression: 0.55 fold) and 4 µg/mL (relative fold expression: 0.59 fold) of TRF (Figure 4b).
Figure 4: Expression of (A) S1004A and (B) mutant P53 (mP53) genes in the wild-type (4T1-WT) and SATB1-silended 4T1 cells treated with TRF (0, 4, 8 or 12 mg/mL) at 37oC for 72 hours in a humidified 5% CO2 incubator was determined using real-time PCR. Results are expressed as relative fold expression of untreated 4T1 cells (Control). *Significantly different from control (p<0.05)
Expressions of β-catenin and E-cadherin
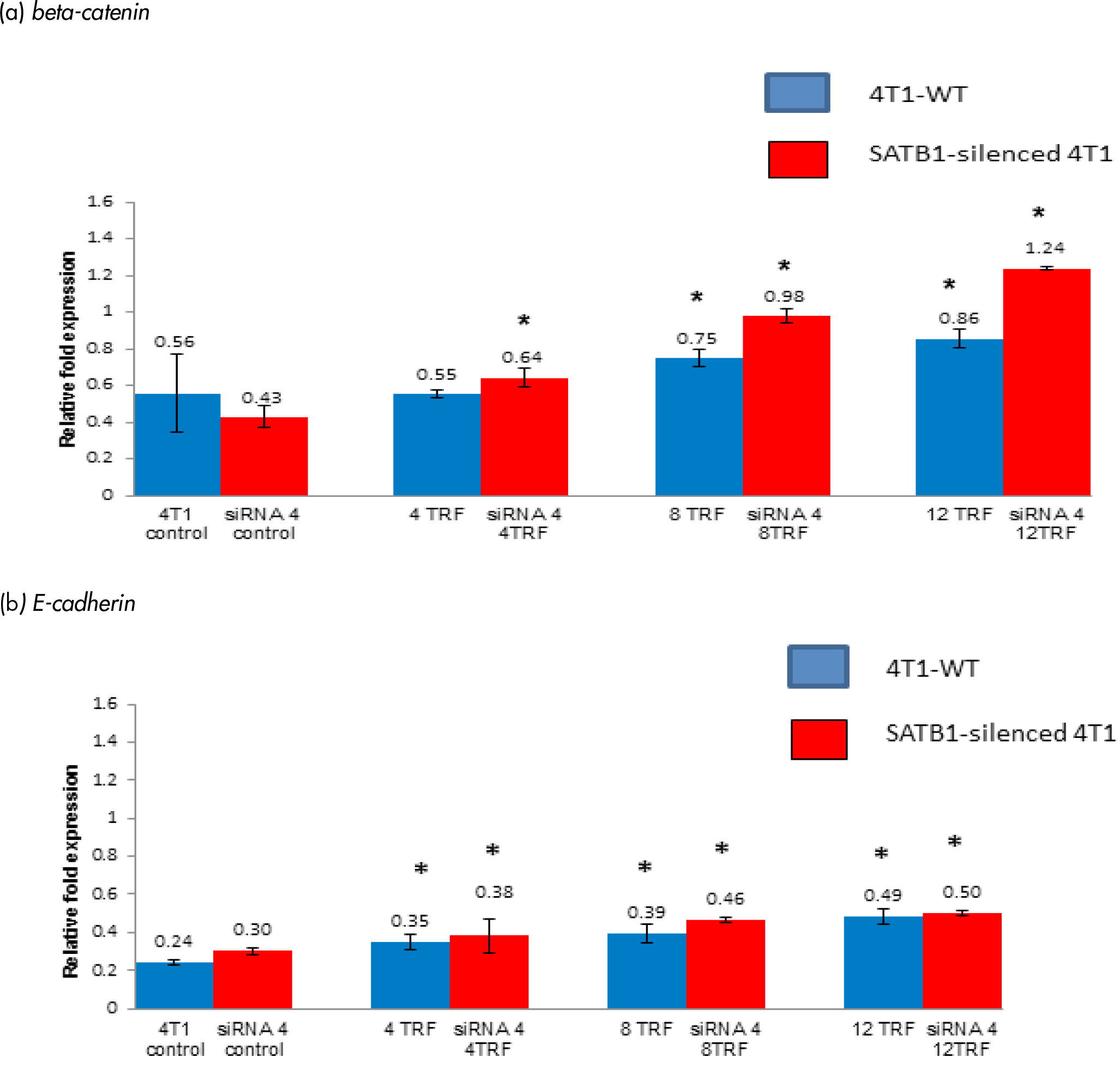
Mutant β-catenin gene has been implicated in the pathogenesis of several cancers [18, 19] and β-catenin was reported to be down-regulated by SATB1 gene (18, 19). The expression of the β-catenin gene was increased in both wild-type and SATB1-silended 4T1 cells treated with different concentrations of TRF (Figure 5a). The expression of the β-catenin gene increased (p<0.05) in SATB1-silenced 4T1 cells when treated with 4 mg/mL (relative fold change: 0.64 fold), 8 mg/mL (relative fold change: 0.98) and 12 mg/mL (relative fold change: 1.24) TRF compared to untreated controls. In the WT 4T1 cells, exposure to TRF also increased (p<0.05) expression of the β-catenin gene [8 mg/mL (relative fold change: 0.75) and 12 mg/mL (relative fold change: 0.86)] of TRF (Figure 5a).
Figure 5: The expression of (a) beta-catenin and (b) E-cadherin genes in the 4T1 wild-type (4T1-WT) and SATB1 gene silenced through the silencing RNA (siRNA) 4T1 approach cultured in the presence or absence of TRF (0, 4, 8 or 12 mg/mL) for 72 hours at 37oC in a humidified 5% CO2 incubator was determined using real time PCR. Results are expressed as relative fold expression of untreated 4T1 cells (Control). *Significantly different from control (p<0.05)
The expression of the E-cadherin gene was increased (p<0.05) in the TRF-treated wild-type and SATB1-silenced 4T1 cells (Figure 5b). Approximately 0.25 relative fold difference was observed when the expression of the E-cadherin gene between control (relative fold change: 0.24) and the same cells exposed to 12 µg/mL TRF (relative fold change: 0.49 fold) while a 0.20-fold difference increase was recorded when compared between the SATB1-silenced untreated 4T1 cells (relative fold change: 0.30 fold) with those that had been exposed to 12 µg/mL TRF (relative fold change: 0.50 fold). Overall, there was an increased E-cadherin in the SATB1-silenced 4T1 cells compared to wild-type cells. Higher expression of the E-cadherin gene was also observed when these cells were exposed to higher concentrations of TRF.
Discussion
Tocotrienol-rich fraction (4 to 12 µg/mL) inhibited proliferation of wild-type (WT) and SATB1-silenced 4T1 cells. The level inhibition observed in the SATB1-silenced 4T1 cells was observed to be greater than the WT cells. The expression of mP53 and S1004A genes were markedly down-regulated in the SATB1-silenced 4T1 cells compared to WT cells.
The mP53 gene contributes to various stages of tumour progressions. This gene is reported to be associated with increased resistance to some of the anti-cancer treatments [17]. In a transgenic mouse model, over-expression of mP53 in certain tissues revealed a variety of manifestations of this gene such as cooperation with additional oncogenic events that accelerate tumour development and make the tumours to be more invasive and aggressive [17].
The S100A4 gene is also known as Mts-1 (metastatin-1), which is a member of the S100 family of calcium-binding proteins [16]. It is a metastasis inducer, making it an important regulator for BC progression [21]. The S1004A gene was also reported to be a SATB1-dependent gene in breast cancer cells [21]. This gene is highly expressed in various metastatic cells and its expression correlates with poor prognosis in some cancer [16, 22 & 23]. Studies with mouse models have shown that the S100A4 gene itself is not tumorigenic, but once a tumour is formed, it enhances tumour invasiveness and metastasis [16, 22 & 23]. The most likely reason for this pattern of expression in the SATB1-silenced 4T1 is because the S1004A gene may be a SATB1-dependent gene in the 4T1 cells. With TRF treatment and/or silencing of the SATB1 gene in the 4T1 cancer cells, we found that these approached can reduce tumour growth as wells as down-regulate the expression of some genes that are responsible for cancer growth and development such as mP53 and S1004A.
Wild-type human BC cells (e.g. MDA-MD 231) are reported to express high levels of the SATB1 gene (16). Studies have shown that the expression of β-catenin and E-cadherin genes are down-regulated in MDA-MD 231 human BC cells due to the expression of the SATB1 gene [20, 21 & 24]. The β-catenin is a member of the canonical Wnt pathway where a network of proteins that are normally involved in embryogenesis and cancer [20, 21 & 24]. E-cadherins are known as a tumour suppressor genes and abnormal expression of this gene has been associated with the development of metastases in BC patients [20]. The finding from the present study corroborates with previous findings that SATB1 plays a role in the inhibiting expression of β-catenin and E-cadherin genes [16, 21] and silencing SATB1 gene appear to increase the expression of both these genes. In addition, exposure to TRF also appears to inhibit SATB1 expression, which in turn increases expression of both of β-catenin and E-cadherin genes.
Conclusion
In order to evaluate the importance of the SATB1 expressions in BC cells, we generated SATB1-silenced 4T1 cells using the siRNA technique. The viability of the SATB1-silenced 4T1 cells decreased when these cells were exposed to TRF. Further analysis showed that in SATB1-silenced 4T1 cells treated with TRF, there was marked down-regulation of the mP53 and S1004A genes, which are responsible for tumour progression as well as up-regulation of the β-catenin and E-cadherin genes.
Author’s Contribution
Conceptualization, A.R.; methodology, A.R and S.C.; software, S.R and S.C.; validation, A.R., S.C. and S.R.; formal analysis, A.R., S.C. and S.R.; investigation, S.R.; resources, A.R., S.C. and S.R.; writing—original draft preparation, S.R.; writing—review and editing, A.R and S.C.; visualization, A.R. and S.R.; supervision, A.R and S.C. All authors have read and agreed to the published version of the manuscript. All authors have read and agreed to the published version of the manuscript.
Funding
This work was supported by grants from Malaysian Palm Oil Board PD119/08 and International Medical University (4.9/5.5/2007).
Acknowledgment
The authors would like to thank the Director General of the Malaysian Palm Oil Board (MPOB) and International Medical University (IMU) for sponsoring this study.
References
- Nachreiner I, Hussain AF, Wullner U, Machuy N, Meyer TF, et al. (2019) Elimination of HER3-expressing breast cancer cells using aptamer-siRNA chimeras. Experimental and therapeutic Medicine 18: 2401–2412. [View]
- Baylin SB (2005) DNA methylation and gene silencing in cancer. Nature Clinical Practice Oncology 2: S4-S11. [View]
- Hannon GJ (2002) RNA interference. Nature 418:244–251. [View]
- Huynh A, Madu CO, Lu Y (2018) siRNA: A Promising New Tool for Future Breast Cancer Therapy. Oncomedicine 3: 74–81. [View]
- Elbashir SM, Harborth J, Lendeckel W, Yalcin, A, Weber K, Tuschl T (2001) Duplexes of 21-nucleotide RNAs mediate RNA interference in cultured mammalian cells. Nature 411: 494–8. [View]
- Lam JK, Chow MY, Zhang Y, Leung SW (2015) siRNA versus miRNA as therapeutics for gene silencing. Molecular therapy. Nucleic acids. 4: e252. [View]
- Abdul Hafid SR, Radhakrishnan AK, Nesaretnam N (2010) Tocotrienols are Good Adjuvants for Developing Cancer Vaccines. BMC (Cancer) Journal 10: 5. [View]
- Abdul Hafid SR, Shahrim Z (2015) The potential of tocotrienols in cancer immunotherapy and wound healing. Palm Oil Development (POD) 61: 14–20. [View]
- Abdul Hafid SR, Chakravarthi S, Nesaretnam K, Radhakrishnan AK (2013) Tocotrienol-Adjuvanted Dendritic Cells Inhibit Tumor Growth and Metastasis: A Murine Model of Breast Cancer. PLoS ONE 8: e74753. [View]
- Abdul Hafid SR, Radhakrishnan AK (2019) Palm Tocotrienol-Adjuvanted Dendritic Cells Decrease Expression of the SATB1 Gene in Murine Breast Cancer Cells and Tissues. Vaccines 7: 198. [View]
- EL Naaman C, Grum-Schwensen B, Mansouri A (2004) Cancer predisposition in mice deficient for the metastasis-associated Mts1(S100A4) gene. Oncogene 23: 3670–3680. [View]
- Eran R Andrechek, William R Hardy, Michael A Laing, William J Muller (2004) Germ-line expression of an oncogenic erbB2 allele confers resistance to erbB2-induced mammary tumorigenesis. Proceedings of the National Academy of Sciences 101: 4984–4989. [View]
- Criswell, Tracy & Arteaga, Carlos (2007) Modulation of NF B Activity and E-cadherin by the Type III Transforming Growth Factor beta Receptor Regulates Cell Growth and Motility. The Journal of biological chemistry 282: 32491–500. [View]
- Rui Zhang, Adrienne, Ma, Stefan J Urbanski, Donna-Marie, McCafferty (2007) Induction of inducible nitric oxide synthase: a protective mechanism in colitis-induced adenocarcinoma. Carcinogenesis 28: 1122–1130. [View]
- Jayashree V, Thenmozhi N (2018) In-vitro anti-proliferative assay and cell viability activity of baicalein using breast cancer cell line. Int J Pharm Sci Res 9: 1620–24. [View]
- Matsuura I, Lai CY, Chiang KN (2010) Functional interaction between Smad3 and S100A4 (metastatin-1) for TGF- -mediated cancer cell invasiveness. Biochemical Journal 426: 327–335. [View]
- Oren M, Rotter V (2010) Mutation P53 gain-off function in cancer. Cold Spring Harb Perspect Biol 2: a001107. [View]
- Patani N, Jiang W, Mansel R, Newbold R, Mokbel K (2009) The mRNA expression of SATB1 and SATB2 in human breast cancer. Cancer Cell International 9: 18. [View]
- Cai S, Han HJ, Kohwi-Shigematsu T (2003) Tissue-specific nuclear architecture and gene expression regulated by SATB1. Nature Genetic 34: 42–51. [View]
- Kowalski P, Rubin MA, Kleer CG (2003) E-cadherin expression in primary carcinomas of the breast and its distant metastases. Breast Cancer Research 5: R217-R222. [View]
- Hans HJ, Russo J, Kohwi Y, Kohwi-shigematsu T (2008) SATB1 reprogremmes gene expression to promote breast tumour growth and metastasis. Nature 452: 187–93. [View]
- Kohwi-Shigematsu T, Mass K, Bode J (1997) A thymocyte factor SATB1 suppresses transcription of stably integrated factor attachment region-linked reporter genes. Biochemistry, 1997; 36: 12005–12010. [View]
- Case SS, Huber P, Lioyd JA (1999) The gamma PE complex contains both SATB1 and HOXB2 and has positive and negative roles in human gamma-globin gene regulation. DNA cell Biology 18: 805–817. [View]
- Kourtidis A,Lu R, Pence L, Anastasiadis PZ (2017) A central role for cadherin signaling in cancer. Experimental Cell Research 2017; 358: 78–85. [View]
Article Type
Research Article
Publication history
Received: October 05, 2025
Accepted: October 15, 2025
Published: October 19, 2025
Citation:
Abd Hafid SR, Radhakrishnan AK, Chakravarthi S (2025) Silencing SATB1 gene Suppressed Expression of mP53, S1004A Genes and Increased Expression of b-catenin and E-cadherin Genes in Murine Mammary Cancer Cells in the Presence and Absence of Tocotrienol-Rich Fraction (TRF). Cancer Stud and Therap I 03(01): 45–54.
Sitti Rahma Abd Hafid1,2,*, Ammu Kutty Radhakrishnan2,3 and Srikumar Chakravarthi4
1Malaysian Palm Oil Board, 6 Persiaran Institusi, Bandar Baru Bangi, 43000 Selangor, Malaysia
2Faculty of Medicine and Health, International Medical University, 126 Jalan 19/155B, Bukit Jalil, 57000 Kuala Lumpur, Malaysia
3Jeffrey Cheah School of Medicine and Health Sciences, Monash University Malaysia, Jalan Lagoon Selatan, 47500 Bandar Sunway, Selangor Darul Ehsan, Malaysia
4Faculty of Medicine, Bioscience and Nursing, MAHSA University, Jln SP2, Bandar Saujana Putra, 42610 Jenjarom, Selangor Darul Ehsan, Malaysia
*Corresponding author
Dr Sitti Rahma Abd Hafid,
Malaysian Palm Oil Board,
6 Persiaran Institusi,
Bandar Baru Bangi,
43000 Selangor,
Malaysia;
Phone: +603-87694566;